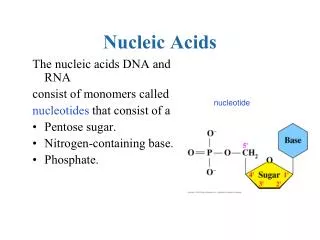
Nucleic Acids: RNA and chemistry
470 likes | 630 Vues
Nucleic Acids: RNA and chemistry. Andy Howard Introductory Biochemistry 13 October 2009. RNA: structure & types mRNA tRNA rRNA Small RNAs. DNA & RNA Hydrolysis alkaline RNA, DNA nucleases Restriction enzymes DNA & RNA dynamics and density measurements. What we’ll discuss.

Nucleic Acids: RNA and chemistry
E N D
Presentation Transcript
Nucleic Acids:RNA and chemistry Andy HowardIntroductory Biochemistry13 October 2009 Biochemistry:Nucleic Acids II
RNA: structure & types mRNA tRNA rRNA Small RNAs DNA & RNA Hydrolysis alkaline RNA, DNA nucleases Restriction enzymes DNA & RNA dynamics and density measurements What we’ll discuss Biochemistry:Nucleic Acids II
Ribonucleic acid • We’re done with DNA for the moment. • Let’s discuss RNA. • RNA is generally, but not always, single-stranded • The regions where localized base-pairing occurs (local double-stranded regions) often are of functional significance Biochemistry:Nucleic Acids II
RNA physics & chemistry • RNA molecules vary widely in size, from a few bases in length up to 10000s of bases • There are several types of RNA found in cells Type % %turn- Size, Partly Role RNA over bases DS? mRNA 3 25 50-104 no protein template tRNA 15 21 55-90 yes aa activation rRNA 80 50 102-104 no transl. catalysis & scaffolding sRNA 2 4 30-103 ? various Biochemistry:Nucleic Acids II
Messenger RNA • mRNA: transcription vehicleDNA 5’-dAdCdCdGdTdAdTdG-3’RNA 3’- U G G C A U A C-5’ • typical protein is ~500 amino acids;3 mRNA bases/aa: 1500 bases (after splicing) • Additional noncoding regions (see later) brings it up to ~4000 bases = 4000*300Da/base=1,200,000 Da • Only about 3% of cellular RNA but instable! Biochemistry:Nucleic Acids II
Relative quantities • Note that we said there wasn’t much mRNA around at any given moment • The amount synthesized is much greater because it has a much shorter lifetime than the others • Ribonucleases act more avidly on it • We need a mechanism for eliminating it because the cell wants to control concentrations of specific proteins Biochemistry:Nucleic Acids II
mRNA processing in Eukaryotes Genomic DNA Unmodified mRNA produced therefrom • # bases (unmodified mRNA) = # base-pairs of DNA in the gene…because that’s how transcription works • BUT the number of bases in the unmodified mRNA > # bases in the final mRNA that actually codes for a protein • SO there needs to be a process for getting rid of the unwanted bases in the mRNA: that’s what splicing is! Biochemistry:Nucleic Acids II
Splicing: quick summary Genomic DNA transcription Unmodified mRNA produced therefrom exon intron exon intron exon intron • Typically the initial eukaryotic message contains roughly twice as many bases as the final processed message • Spliceosome is the nuclear machine (snRNAs + protein) in which the introns are removed and the exons are spliced together splicing exon exon exon translation (Mature transcript) Biochemistry:Nucleic Acids II
Heterogeneity via spliceosomal flexibility • Specific RNA sequences in the initial mRNA signal where to start and stop each intron, but with some flexibility • That flexibility enables a single gene to code for multiple mature RNAs and therefore multiple proteins Biochemistry:Nucleic Acids II
Transfer RNA • tRNA: tool for engineering protein synthesis at the ribosome • Each type of amino acid has its own tRNA, responsible for positioning the correct aa into the growing protein • Roughly T-shaped or Y-shaped molecules; generally 55-90 bases long • 15% of cellular RNA Phe tRNAPDB 1EVV76 basesyeast Biochemistry:Nucleic Acids II
Secondary and Tertiary Structure of tRNA • Extensive H-bonding creates four double helical domains, three capped by loops, one by a stem • Only one tRNA structure (alone) is known • Phenylalanine tRNA is "L-shaped" • Many non-canonical base pairs found in tRNA Biochemistry:Nucleic Acids II
tRNA structure: overview Biochemistry:Nucleic Acids II
Amino acid linkage to acceptor stem Amino acids are linked to the 3'-OH end of tRNA molecules by an ester bond formed between the carboxyl group of the amino acid and the 3'-OH of the terminal ribose of the tRNA. Biochemistry:Nucleic Acids II
Yeast ala-tRNA • Note nonstandard bases and cloverleaf structure Biochemistry:Nucleic Acids II
Ribosomal RNA • rRNA: catalyic and scaffolding functions within the ribosome • Responsible for ligation of new amino acid (carried by tRNA) onto growing protein chain • Can be large: mostly 500-3000 bases • a few are smaller (150 bases) • Very abundant: 80% of cellular RNA • Relatively slow turnover 23S rRNAPDB 1FFZ602 basesHaloarcula marismortui Biochemistry:Nucleic Acids II
Ribosomal composition (fig.10.22) • Bacterial ribosome • 30S subunit: 16S RNA + 21 proteins • 50S subunit:23S RNA + 5S RNA + 31 proteins • Eukaryotic ribosome • 40S subunit: 18S RNA + 33 proteins • 60S subunit:(28S+5.85S) RNA, 5S RNA + 49 proteins Biochemistry:Nucleic Acids II
Small RNA • sRNA: few bases / molecule • often found in nucleus; thus it’s often called small nuclear RNA, snRNA • Involved in various functions, including processing of mRNA in the spliceosome • Some are catalytic • Typically 20-1000 bases • Not terribly plentiful: ~2 % of total RNA Protein Prp31complexed to U4 snRNAPDB 2OZB33 bases + 85kDa heterotetramerHuman Biochemistry:Nucleic Acids II
siRNAs and gene silencing • Small interfering RNAs block specific protein production by base-pairing to complementary seqs of mRNA to form dsRNA • DS regions get degraded & removed • This is a form of gene silencing or RNA interference • RNAi also changes chromatin structure and has long-range influences on expression Viral p19 protein complexed to human 19-base siRNA PDB 1R9F1.95Å 17kDa protein Biochemistry:Nucleic Acids II
Other small RNAs • 21-28 nucleotides • Target RNA or DNA through complementary base-pairing • Several types, based on function: • Small interfering RNAs (q.v.) • microRNA: control developmental timing • Small nucleolar RNA: catalysts that (among other things) create the oddball bases snoRNA77courtesy Wikipedia Biochemistry:Nucleic Acids II
How many varieties of each class? • mRNA: thousands(one per protein transcript) • tRNA: one per codon plus a few more • rRNA: a few per organism—see rRNA slide • sRNA: dozens (?) Biochemistry:Nucleic Acids II
Unusual bases in RNA • mRNA, sRNA mostly ACGU • rRNA, tRNA have some odd ones Biochemistry:Nucleic Acids II
iClicker quiz • 1. Shown is the lactim form of which nucleic acid base? • Uracil • Guanine • Adenine • Thymine • None of the above Biochemistry:Nucleic Acids II
iClicker quiz #2 • Suppose someone reports that he has characterized the genomic DNA of an organism as having 29% A and 22% T. How would you respond? • (a) That’s a reasonable result • (b) This result is unlikely because [A] ~ [T] in duplex DNA • (c) That’s plausible if it’s a bacterium, but not if it’s a eukaryote • (d) none of the above Biochemistry:Nucleic Acids II
Do the differences between RNA and DNA matter? Yes! • DNA has deoxythymidine, RNA has uridine: • cytidine spontaneously degrades to uridine • dC spontaneously degrades to dU • The only dU found in DNA is there because of degradation: dT goes with dA • So when a cell finds dU in its DNA, it knows it should replace it with dC or else synthesize dG opposite the dU instead of dA Biochemistry:Nucleic Acids II
Ribose vs. deoxyribose • Presence of -OH on 2’ position makes the 3’ position in RNA more susceptible to nonenzymatic cleavage than the 3’ in DNA • The ribose vs. deoxyribose distinction also influences enzymatic degradation of nucleic acids • I can carry DNA in my shirt pocket, but not RNA Biochemistry:Nucleic Acids II
Backbone hydrolysis of nucleic acids in base(fig. 10.29) • Nonenzymatic hydrolysis in base occurs with RNA but not DNA, as just mentioned • Reason: in base, RNA can form a specific 5-membered cyclic structure involving both 3’ and 2’ oxygens • When this reopens, the backbone is cleaved and you’re left with a mixture of 2’- and 3’-NMPs Biochemistry:Nucleic Acids II
Why alkaline hydrolysis works • Cyclic phosphate intermediate stabilizes cleavage product Biochemistry:Nucleic Acids II
The cyclic intermediate • Hydroxyl or water can attack five-membered P-containing ring on either side and leave the –OP on 2’ or on 3’. Biochemistry:Nucleic Acids II
Consequences • So RNA is considerably less stable compared to DNA, owing to the formation of this cyclic phosphate intermediate • DNA can’t form this because it doesn’t have a 2’ hydroxyl • In fact, deoxyribose has no free hydroxyls! Biochemistry:Nucleic Acids II
Enzymatic cleavage of oligo- and polynucleotides • Enzymes are phosphodiesterases • Could happen on either side of the P • 3’ cleavage is a-site; 5’ is b-site. • Endonucleases cleave somewhere on the interior of an oligo- or polynucleotide • Exonucleases cleave off the terminal nucleotide Biochemistry:Nucleic Acids II
An a-specific exonuclease Biochemistry:Nucleic Acids II
A b-specific exonuclease Biochemistry:Nucleic Acids II
Specificity in nucleases • Some cleave only RNA, others only DNA, some both • Often a preference for a specific base or even a particular 4-8 nucleotide sequence (restriction endonucleases) • These can be used as lab tools, but they evolved for internal reasons Biochemistry:Nucleic Acids II
Enzymatic RNA hydrolysis • Ribonucleases operate through a similar 5-membered ring intermediate: see fig. 19.29 for bovine RNAse A: • His-119 donates proton to 3’-OP • His-12 accepts proton from 2’-OH • Cyclic intermediate forms with cleavage below the phosphate • Ring collapses, His-12 returns proton to 2’-OH, bases restored PDB 1KF813.6 kDa monomer bovine Biochemistry:Nucleic Acids II
Variety of nucleases Biochemistry:Nucleic Acids II
Restriction endonucleases • Evolve in bacteria as antiviral tools • “Restriction” because they restrict the incorporation of foreign DNA into the bacterial chromosome • Recognize and bind to specific palindromic DNA sequences and cleave them • Self-cleavage avoided by methylation • Types I, II, III: II is most important • I and III have inherent methylase activity; II has methylase activity in an attendant enzyme Biochemistry:Nucleic Acids II
What do we mean by palindromic? • In ordinary language, it means a phrase that reads the same forward and back: • Madam, I’m Adam. (Genesis 3:20) • Eve, man, am Eve. • Sex at noon taxes. • Able was I ere I saw Elba. (Napoleon) • A man, a plan, a canal: Panama! (T. Roosevelt) • With DNA it means the double-stranded sequence is identical on both strands Biochemistry:Nucleic Acids II
Quirky math question to ponder • Numbers can be palindromic:484, 1331, 727, 595… • Some numbers that are palindromic have squares that are palindromic…222 = 484, 1212 = 14641, . . . • Question: if a number is perfect square and a palindrome, is its square root a palindrome? (answer will be given orally) Biochemistry:Nucleic Acids II
Palindromic DNA • G-A-A-T-T-C • Single strand isn’t symmetric: but the combination with the complementary strand is: • G-A-A-T-T-CC-T-T-A-A-G • These kinds of sequences are the recognition sites for restriction endonucleases. This particular hexanucleotide is the recognition sequence for EcoRI. Biochemistry:Nucleic Acids II
Cleavage by restriction endonucleases • Breaks can be • cohesive (if they’re off-center within the sequence) or • non-cohesive (blunt) (if they’re at the center) • EcoRI leaves staggered 5’-termini: cleaves between initial G and A • PstI cleaves CTGCAG between A and G, so it leaves staggered 3’-termini • BalI cleaves TGGCCA in the middle: blunt! Biochemistry:Nucleic Acids II
iClicker question 3: • 3. Which of the following is a potential restriction site? • (a) ACTTCA • (b) AGCGCT • (c) TGGCCT • (d) AACCGG • (e) none of the above. Biochemistry:Nucleic Acids II
Example for EcoRI • 5’-N-N-N-N-G-A-A-T-T-C-N-N-N-N-3’3’-N-N-N-N-C-T-T-A-A-G-N-N-N-N-5’ • Cleaves G-A on top, A-G on bottom: • 5’-N-N-N-N-GA-A-T-T-C-N-N-N-N-3’3’-N-N-N-N-C-T-T-A-AG-N-N-N-N-5’ • Protruding 5’ ends:5’-N-N-N-N-GA-A-T-T-C-N-N-N-N-3’3’-N-N-N-N-C-T-T-A-AG-N-N-N-N-5’ Biochemistry:Nucleic Acids II
How often? • 4 types of bases • So a recognition site that is 4 bases long will occur once every 44 = 256 bases on either strand, on average • 6-base site: every 46= 4096 bases, which is roughly one gene’s worth Biochemistry:Nucleic Acids II
EcoRI structure • Dimeric structure enables recognition of palindromic sequence • sandwich in each monomer EcoRI pre-recognition complex PDB 1CL8 57 kDa dimer + DNA Biochemistry:Nucleic Acids II
Methylases HhaI methyltransferasePDB 1SVU2.66Å; 72 kDa dimer • A typical bacterium protects its own DNA against cleavage by its restriction endonucleases by methylating a base in the restriction site • Methylating agent is generally S-adenosylmethionine Structure courtesy steve.gb.com Biochemistry:Nucleic Acids II
The biology problem • How does the bacterium mark its own DNA so that it does replicate its own DNA but not the foreign DNA? • Answer: by methylating specific bases in its DNA prior to replication • Unmethylated DNA from foreign source gets cleaved by restriction endonuclease • Only the methylated DNA survives to be replicated • Most methylations are of A & G,but sometimes C gets it too Biochemistry:Nucleic Acids II
How this works • When an unmethylated specific sequence appears in the DNA, the enzyme cleaves it • When the corresponding methylated sequence appears, it doesn’t get cleaved and remains available for replication • The restriction endonucleases only bind to palindromic sequences Biochemistry:Nucleic Acids II