An Introduction to Genome Classification, Analysis and Annotation
250 likes | 548 Vues
An Introduction to Genome Classification, Analysis and Annotation. Genome Analysis: From Sequencing to Annotation. The main feature of genome browsers Is to display all of the available information along a genomic region of interest. Genomic regions contain 4 levels of resolution

An Introduction to Genome Classification, Analysis and Annotation
E N D
Presentation Transcript
An Introduction to Genome Classification, Analysis and Annotation
Genome Analysis: From Sequencing to Annotation • The main feature of genome browsers • Is to display all of the available information along a genomic region of interest. • Genomic regions contain 4 levels of resolution • From the chromosomal level to individual base-pairs.
Genome Analysis • Whole genome analysis • Gene count • Gene classification • Repeat content • Chromosomal duplications • Multi-Genome Analysis • Synteny • Sequence similarity • Gene classification comparisons • Whole genome analysis • Gene count • Gene classification • Repeat content • Chromosomal duplications • Multi-Genome Analysis • Synteny • Sequence similarity • Gene classification comparisons
Gene Count-How do we find genes in genomic sequences? • Gene Predictions: • Fgenesh (http://www.softberry.com) • GenemarkHMM (http://opal.biology.gatech.edu/GeneMark/eukhmm.cgi) • GenScan (http://genes.mit.edu/GENSCAN.html) • Grail (http://compbio.ornl.gov/Grail-1.3/) • Glimmer (http://www.tigr.org/softlab/glimmer/glimmer.html) • Map cDNA sequences to a genome • Sim4 (http://pbil.univ-lyon1.fr/sim4.html) • EST2Genome (http://bioweb.pasteur.fr/seqanal/interfaces/est2genome.html) • Genomewise • BLAT • Homology • blastx
Gene Prediction Types • Known: cDNA evidence/homology • Putative: Gene prediction which has homology to known gene • Unknown: EST matching a gene prediction • Hypothetical: Gene prediction(s) only • Known: cDNA evidence/homology • Putative: Gene prediction which has homology to known gene • Unknown: EST matching a gene prediction • Hypothetical: Gene prediction(s) only
Review: cDNAs and ESTs • Complementary DNA (cDNA)s • Are DNA synthesized from a mature (fully spliced/edited) RNA template • Expressed Sequence Tags (ESTs) • Are short sub-sequences of a transcribed, spliced sequence • May be used to identify gene transcripts • Are instrumental in gene discovery and sequence determination
cDNAs and ESTs reveal exons or gene ends in genome searches Figure 13-10
Using Expressed Sequence Tags(ie. sequencing parts of the processed genes) Advantages • Inexpensive • “Know” sequence is coding • Information about tissue or developmental stage expression Disadvantages • Coverage is incomplete • Position of sequence in the genome is unknown • Only partial information about each gene • No information about structural elements
Gene Prediction Types • Known: cDNA evidence/homology • Putative: Gene prediction which has homology to known gene • Unknown: EST matching a gene prediction • Hypothetical: Gene prediction(s) only • Known: cDNA evidence/homology • Putative: Gene prediction which has homology to known gene • Unknown: EST matching a gene prediction • Hypothetical: Gene prediction(s) only
Gene Prediction using Homology • BLAST as a tool to identify regions of sequence similarity in other organisms as compared to a query sequence • Example: In gene finding, the search tool BLASTX translates a nucleotide query sequence in all six reading frames to search protein databases.
Gene Prediction Types • Known: cDNA evidence/homology • Putative: Gene prediction which has homology to known gene • Unknown: EST matching a gene prediction • Hypothetical: Gene prediction(s) only • Known: cDNA evidence/homology • Putative: Gene prediction which has homology to known gene • Unknown: EST matching a gene prediction • Hypothetical: Gene prediction(s) only
The information content of the genome includes binding sites Figure 13-9
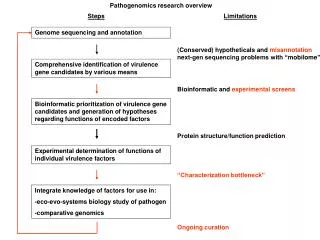
Genome searches hunt for various binding sites • The different forms of evidence • Include cDNAs, ESTs, BLAST similarity hits, codon bias, and motif hits. • Are integrated to make gene predictions . • Where multiple classes of evidence are found to be associated with a particular genomic DNA sequence • There is a greater confidence in the likelihood that a gene prediction is accurate. Figure 13-11
Summary: many forms of evidence are integrated to make gene predictions Figure 13-12
Gene Classification • Automated • Similarity search against an annotated database • Swiss-Prot (http://www.expasy.ch/sprot/) • Nr • Protein Domain search • i. Pfam (http://www.sanger.ac.uk/Software/Pfam/) • ii. Prosite • iii. Prints • iv. Prodom • v. Interpro (http://www.ebi.ac.uk/interpro/scan.html) • 2) Curated • Similar to above but usually people will verify results through literature searches
Looking for Repeats • RepeatMasker can find and mask repeats in DNA sequence • RepeatMasker can be found at http://woody.embl-heidelberg.de/repeatmask/ or http://repeatmasker.genome.washington.edu/ • 3. RepeatMasker is often run on genomic sequences before doing gene predictions
Genomes on Display • UCSC Browser • Ensembl browser • NCBI Browser • GMOD
Genome Analysis • Whole genome analysis • Gene count • Gene classification • Repeat content • Chromosomal duplications • Multi-Genome Analysis • Synteny • Sequence similarity • Gene classification comparisons • Whole genome analysis • Gene count • Gene classification • Repeat content • Chromosomal duplications • Multi-Genome Analysis • Synteny • Sequence similarity • Gene classification comparisons
The mouse and human genome • have large syntenic blocks of genes in common