Protein Kinase Structure and Function Introduction
Protein Kinase Structure and Function Introduction 500 protein kinases encoded in the human genome 1.5 % of our genetic diversity Protein kinases regulate at some level almost all cellular processes. Cellular proliferation Cell cycle Replication Translation …..


Protein Kinase Structure and Function Introduction
E N D
Presentation Transcript
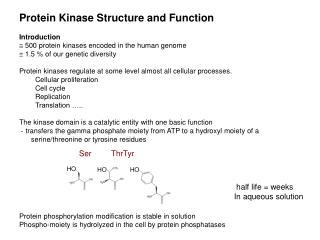
Protein Kinase Structure and Function • Introduction • 500 protein kinases encoded in the human genome • 1.5 % of our genetic diversity • Protein kinases regulate at some level almost all cellular processes. • Cellular proliferation • Cell cycle • Replication • Translation ….. • The kinase domain is a catalytic entity with one basic function • - transfers the gamma phosphate moiety from ATP to a hydroxyl moiety of a serine/threonine or tyrosine residues • Protein phosphorylation modification is stable in solution • Phospho-moiety is hydrolyzed in the cell by protein phosphatases Ser Thr Tyr HO HO HO half life = weeks In aqueous solution
Role of protein phosphorylation: Phosphorylation induces a conformational change or generates a protein interaction ligand. i. Conformational change: Phosphorylation of a residue causes a gross conformational change in target protein that affects function. -e.g. protein kinases themselves (very tight ON/OFF switch) Insulin receptor kinase ii. Promote complex formation by the generation of protein interaction ligands: -phosphorylation generates a product that is recognized (or no longer recognized) by a protein interaction module. This can serve to define networks of interacting proteins. -SH2 domain (src homology 2) Src SH2 binds pYEEI ligands -PTB domain (phosphotyrosine binding domain) Shc and IRS-1 NPXpY ligands -14,3,3 domain (phosphoserine binding domains) XXXpSerXXX ligands -FHA domain (phosphothreonine binding domains) XXXpThrXXX ligands -WD40 repeat domain, cdc4 b-trcp – diphophopeptide motif pThrPXXpThr
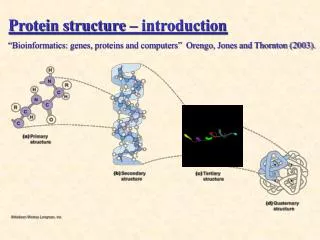
SH2 Domain: The prototypical phospho recognition module • The prototype ‘interaction module’ discovered and characterized by Dr. Tony Pawson • -100 amino acid module consists of a central b-sheet of (4 to 6 b-strands) and two a-helices • -binding site lies across the sheet structure flanked by the two helices • -recognizes phospho-tyrosine containing motifs (reads out sequence Yxxx after phospho-tyrosine) • - SH2 domains bind the phosphotyrosine using a conserved mechanism • - Single Arg mutation disables all phospho-peptide binding –surgical for dissecting biological function • The phosphotyrosine binding site is most highly conserved part of SH2 domains • Invariant arginine and deep pocket are most critical binding features Y 416 Y 527
The eukaryotic protein kinase domain 12 highly conserved motifs Structural elements of the kinase domain N-lobe –b sheet + one helix aC C-lobe –mainly a-helical Cleft region between lobes -nucleotide binding -location of phospho acceptor site -catalytic residues -activation segment (blue) C-lobe = phospho-acceptor binding site
35kDa 300a.a. core -12 highly conserved motifs termed sub domains -12 near invariant residues PKA = prototype structure (Susan Taylor Lab) Hundreds of structures now available in PDB N-lobe Sub-domain function 1 glycine rich loop flexible flap that tethers sugar and non- hydrolyzable phosphate groups 2 invariant Lys glu-lys salt bridge orients a & b 3 invariant Glu phosphate 5 hinge region coordinates adenine ring of ATP 6 invariant Asp catalytic base in catalytic loop (HRDxxxxN) removes proton from target hydroxyl site (Asn orients Asp) 7 invariant Asp binds mg2+ that in turn binds and (DFG) orients b & g phosphates of ATP 7.5 Activation regions spanning DFG and APE segment motif usually contains site of phophoregulation 8 Helix aEF Determines Ser/Thr vs Tyrosine (APE) specificity and flanking sequences (P+1 loop) D5 aEF C-lobe
Catalytic mechanism is well understood Catalytic mechanism is conserved for all kinases
All eukaryotic protein kinases transfer phosphate the same way and look very much alike in their active states • How do the 500 kinases differ to allow each to regulate specific aspects of biology? • Origin of diversity lies with: • Catalytic switching mechanisms • -the ability to switch on and off in response to specific upstream signals • Substrate targeting mechanisms • The ability to select a restricted set of substrates in the cell any one time • -15,000+ proteins in the genome • - each with many phosphorylatable sites (ser/thr, tyr)
Features of the kinase that facilitate diversification of function i. the protein kinase catalytic domain is structurally pliant / flexible bilobal nature – connection by flexible hinge N-terminal sheet – easily deformed aC helix –position easily modulated g-loop – gly is inherently flexible activation segment ii. strict conformational requirements for catalysis – perturb structure of any conserved element and kinase is inactivated iii. extensive surface provides opportunities for the evolution of secondary peptide binding sites to complement the peptide binding site at the catalytic site -although core catalytic structure is conserved exposed surfaces are highly variable iv. modular – kinase domain is commonly linked to non-catalytic interaction domains (eg. SH2 PTB, …) These factors provide many opportunities for diversification of substrate recognition and regulation and indeed diverse mechanisms of regulation have been uncovered and many more remain to be solved. We will survey paradigms uncovered by x-ray crystallographic methods.
Substrate targeting mechanisms Active site directed specificity i. Ser/Thr versus Tyrosine ii. Proline directed iii. Phospho-priming P+1 loop of activation segment defines Ser/Thr versus Tyr preference - serves as a platform for main chain of substrate Ser Thr Tyr Held close Held far What about dual specificity kinases like PKR?
Tyrosine kinases Tyr Serine/Threonine kinases Ser/Thr versus Tyrosine kinases (easy to predict)
Unique highly constrained back bone conformation necessitates specialized infrastructure MAP kinases cyclin dependent Protein kinases PROLINE (S/T-P) DIRECTED protein kinases (easy to predict)
SRPK GSK3a,b Phospho-priming dependent protein kinases variable mechanisms make prediction difficult
Higher order Substrate Targeting Mechanisms (augmenting specificity of the active site) Secondary Peptide Docking Sites Highly specific to subfamilies hypothetical substrate Acceptor site motif Docking site motif C N OH Very versatile and pervasive but hard to predict substrate-kinase relationship due to variation in placement and degenerate nature of the motifs Substrates may not look anything like each other except for presence of two short motifs Each kinase can / has evolved tens to hundreds of substrates
MAP kinases Cyclin dependent protein kinases Mechanisms are conserved across closely related subfamilies
Higher order Substrate Targeting Mechanisms – cont. Best characterized example = PKR (an eIF2a kinase) eIF2a Substrate targeting based on Domain - Domain recognition -Highly specialized -Relatively monogamous
Diverse Sensing Common Target Substrate recognition by the eIF2a protein Kinases heme deficiency HRI viral invasion P Ser51 PKR Cellular Stress eIF2a amino acid starvation GCN2 Inhibition of Protein Synthesis misfolded proteins PERK distinct regulatory domains similar catalytic domains
Molecular basis for specificity revealed by x-ray crystallography eIF2a PKR Specificity arise from non-cononical aG helix – very diagnostic
Second example of domain based substrate targeting Type I TGF-b receptor kinases (7 members) / SMAD substrates (5 members) Globular MH2 domain - TGFB family kinases phoshorylate Smad family of proteins and almost nothing else - Presence of MH2 domain in Smad proteins very diagnostic - Recognition is phosphorylation dependent Acceptor site C-terminal to MH2 domain Structure of the complex has not yet been determined
TGF-b receptor family • eIF2a kinase family • PKR • GCN2 • PERK • HRI Domain-Domain dependent substrate recognition
Y 416 Y 527 Substrate targeting mechanisms Use of interaction domains Splicing modules together is easier than evolving entirely new interaction surfaces Very versatile Same issues as kinases using peptide docking sites SH3 SH2 Hypothetical substrate Docking motif 1 Acceptor site motif C N xYxxx pxxp pYEEI Docking motif 2