1.) DNA Extraction
300 likes | 348 Vues
1.) DNA Extraction. Follow Kit Grind sample Mix with solution and spin Bind, Wash, Elute. Is DNA present?. Gel Electrophoresis DNA has negative charge. In Vivo DNA unwound (denatured) by enzymes RNA polymerase makes “primer” DNA polymerase adds nucleotides. In Vitro

1.) DNA Extraction
E N D
Presentation Transcript
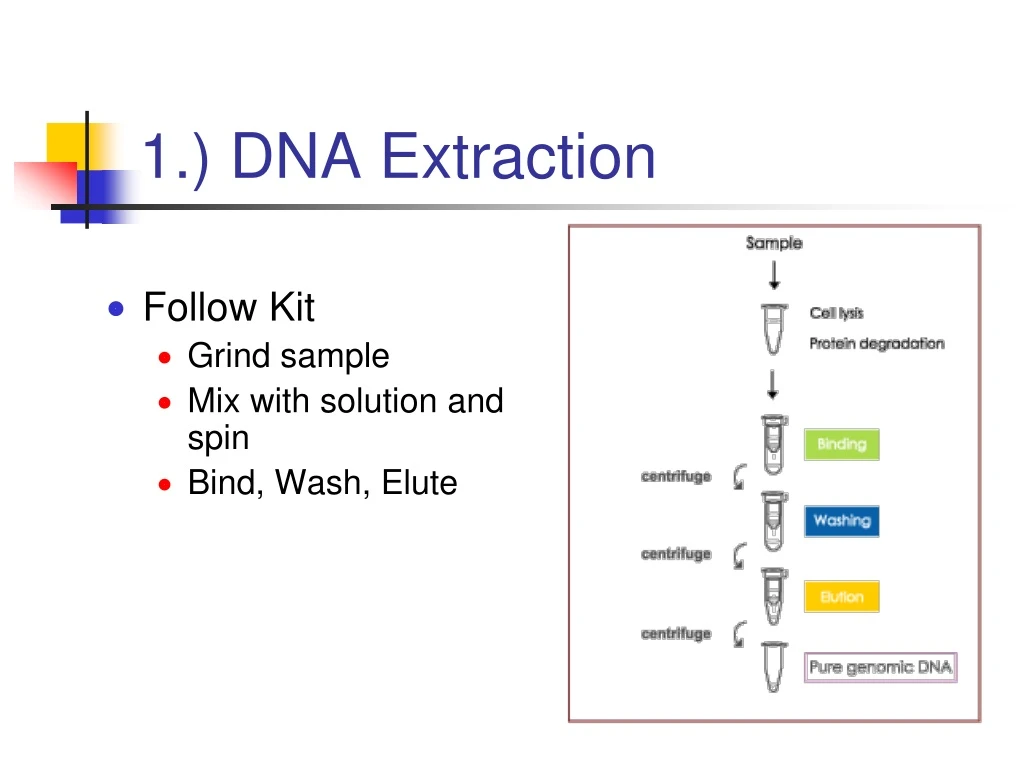
1.) DNA Extraction • Follow Kit • Grind sample • Mix with solution and spin • Bind, Wash, Elute
Is DNA present? • Gel Electrophoresis • DNA has negative charge
In Vivo DNA unwound (denatured) by enzymes RNA polymerase makes “primer” DNA polymerase adds nucleotides In Vitro DNA unwound by heat Primers are added DNA polymerase adds nucleotides DNA Replication
PCR Protocol • Primer I (reverse) • Primer II ( forward) • DNA template • Taq DNA polymerase • MgCl2 • Deoxyribonucleotide triphosphate (dNTP) • PCR buffer • Deionized water
2) Polymerase Chain Reaction • Amplify certain sequence into billionfold • Put DNA sample into tube and mix with ingredients: • Stages of PCR process • Denature • Anneal • Extension
5’ 3’ 3’ 5’ 5’ 3’ 2) Annealing Forward Primer Reverse Primer 3’ 5’ 5’ 3’ 5’ 3’ 5’ 3’ 3) Extension Add dNTP’s 3’ 5’ 5’ 3’ 5’ 3’ 1)Denaturation “Unwind Helix”
Cycle Sequencing • Same as PCR • Denature • Anneal (forward and reverse primers in separate reactions) • Extension • dNTP • ddNTP • Different size of DNA fragments
ddNTP • a dideoxyribonucleotide triphosphate (ddNTP), share the same structure as a normal dNTP, with the exception of the 3' OH group, which is replaced by an H:
dNTP OH
“ddNTP” H
…ddNTP • Dideoxynucleotides, or ddNTPs, are nucleotides lacking a 3'-hydroxyl (-OH) group on their deoxyribose sugar. • Since deoxyribose already lacks a 2'-OH, dideoxyribose lacks hydroxyl groups at both its 2' and 3' carbons. • The lack of this hydroxyl group means that, after being added by a DNA polymerase to a growing nucleotide chain, no further nucleotides can be added as no phosphodiester bond can be created based on the fact that deoxyribonucleoside triphosphates (which are the building blocks of DNA) allow DNA chain synthesis to occur through a condensation reaction between the 5' phosphate (following the cleavage of pyrophospate) of the current nucleotide with the 3' hydroxyl group of the previous nucleotide.
…ddNTP • The dideoxyribonucleotides do not have a 3' hydroxyl group, hence no further chain elongation can occur once this dideoxynucleotide is on the chain. • This can lead to the determination of the DNA sequence. Thus, these molecules form the basis of the dideoxy chain-termination method of DNA sequencing, which was developed by Frederick Sanger (1977).
dNTP & ddNTP • Four separate reaction tubes are required, each one containing radioactively labeled DNA primers, DNA Polymerase II, and an ample amount of all 4 dNTP (dATP, dTTP, dGTP, dCTP) each to be integrated into the DNA strand being synthesized as a nucleotide. • Each reaction tubes contain a different ddNTP, allowing each tube to identify a different nucleotide along the strand. • For example, one tube would contain a ddATP, enabling that reaction tube to identify all the A's being integrated into the synthesizing strand • Thus all the T's in the complementary template strand (recall that T nucleotides are complementary and base pair with A nucleotides). • The above is required for the rest of the tubes with dTTP, dGTP, dCTP
dNTP & ddNTP • All 4 dNTPs and a different ddNTP are added to each reaction tube in a ratio of around 300:1, and Polymerase will randomly integrate either a dNTP or a ddNTP into the synthesizing strand if the ddNTP complements with the nucleotide on the template strand.
Sequencing method • Before the DNA can be sequenced, it has to be denatured into single strands using heat. • Next a primer is annealed to one of the template strands. This primer is specifically constructed so that its 3' end is located next to the DNA sequence of interest. • Either this primer or one of the nucleotides should be radioactively or fluorescently labeled so that the final product can be detected on a gel (Russell, 2002). • Once the primer is attached to the DNA, the solution is divided into four tubes labeled "G", "A", "T" and "C". Then reagents are added to these samples as follows:
….Sequencing method Recap • First, anneal the primer to the DNA template (usually single stranded), Example: • 5'-GAATGTCCTTTCTCTAAG 3'-GGAGACTTACAGGAAAGAGATTCAGGATTCAGGAGGCCTACCATGAAGATCAAG-5' • Then split the sample into four aliquots, in tubes labeled "G", "A", "T" and "C" and add the following substrates to the respective tubes: • "G" tube: all four dNTP's, ddGTP and DNA polymerase • "A" tube: all four dNTP's, ddATP and DNA polymerase • "T" tube: all four dNTP's, ddTTP and DNA polymerase • "C" tube: all four dNTP's, ddCTP and DNA polymerase
…. Sequencing method • When a polymerase (e.g. Klenow fragment) is added to the tubes, the synthetic reaction proceeds until, by chance, a dideoxynucleotide is incorporated instead of a deoxynucleotide. • This is a "chain termination" event, because there is a 3' H instead of a 3' OH group. • Since the synthesized DNA contains some radiolabeled (or chemically labeled) substrates, the products can be detected and distinguished from the template.
…Sequencing method • As the DNA is synthesized, nucleotides are added on to the growing chain by the DNA polymerase. • However, on occasion a dideoxynucleotide is incorporated into the chain in place of a normal nucleotide, which results in a chain-terminating event. • For example if we looked at only the "G" tube, we might find a mixture of the following products:
Figure 1: An example of the potential fragments that could be produced in the "G" tube. The fragments are all different lengths due to the random integration of the ddGTP's (Metzenberg).
ddNTP = Array of Fragments • http://www.dnalc.org/ddnalc/resources/cycseq.html
Cycle Sequencing Results • Electropherogram - laser reads DNA fragment • Adenine • Thymine • Cytosine • Guanine
References • Introduction to Biotechnology by W.J. Thieman and M.A. Palladino. Pearson & Benjamin Cummings 2nd edition. • http://en.wikipedia.org • http://www.ocf.berkeley.edu/~edy/genome/sequencing.html • http://www.escience.ws/b572/L8/L8.htm • http://www.bio.davidson.edu/Courses/Molbio/MolStudents/spring2003/Obenrader/sanger_method_page.htm • PPT lecture materials- Courtesy DRs T Mc Elroy & PN Achar