Biofilms
Biofilms. Microbial Biochemistry. Definition of a Biofilm. Biofilms are communities of microorganisms in a matrix that joins them together and to living or inert substrates

Biofilms
E N D
Presentation Transcript
Biofilms Microbial Biochemistry
Definition of a Biofilm • Biofilms are communities of microorganisms in a matrix that joins them together and to living or inert substrates • Biofilms are surface-attached communities of bacteria, encased in an extracellular matrix of secreted proteins, carbohydrates, and/or DNA, that assume phenotypes distinct from those of planktonic cells
Formation of biofilms in nature • Biofilms offer their member cells several benefits • Biofilms are diverse from their formation on teeth as plaques and submerged rocks in a stream
Biofilms BIOFILMS may form: • On solid substratums in contact with moisture • On soft tissue surfaces in living organisms • At liquid-air interfaces.
Study of biofilms • Formation of multicellular communities depends on the production of extracellular substances( matrix) • Diversity in the formation of the matrix
Pathogens that have been studied for the formation of biofilms • Staphylococcus aureus • Staphylococcus mutans • Salmonella typhi • Enterococcus faecalis • Pseudomonas aeruginosa
Biofilms – Now a Universal Feature • Now scientists believe biofilm formation is a universal feature of all bacteria
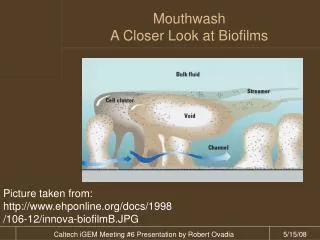
Biofilm characteristics • Submerged biofilms seems to form columns and mushroom like projections that are separated by water-filled channels • Floating biofilms form a skin or pellicle at the air- liquid interface – shows organization of cells with the matrix at the outside • Films that form on the surface of solid media such as agar or other surfaces
Steps in biofilm formation • Initiation of biofilm formation – interaction of cells with a surface or with each other • Films aggregate • Then the cells form an extracellular matrix • Structure of biofilms are dramatically different due to the specific organisms in the film and environmental conditions
Figure from: Kolter, R. and R. Losick. 1998. One for all and all for one. Science 280:226-227.
Exopolysaccharides • Exopolysaccharides • In the glycoclyx contribute to biofilm formation
Matrix • Key components of the matrix are extracellular polysaccharides and proteins • Dead cells have also been identified in biofilms • Extracellular DNA is also important
Polysaccharides • Carbohydrates significantly impact bacterial virulence • Bacteria have capsular polysaccharides and exopolysaccharides • The polysaccharides are not soluble and do not disassociate with the bacterial cells
Polysaccahrides • Many bacteria have been found to produce cellulose • This is a novel finding in the case of Salmonella typhimurium and E. coli • The bacterium Gluconacetobacter xylinus has been recognized as a cellulose producer • Many other bacteria have genes homologous to the bcs, bacterial cellulose synthesis genes • Vibrio cholera does not appear to have a gene which encodes a cellulose – but the bacterium has two domains homologous to Gluconeacetobacter.
Biofilm Genetics - initiation • Staphylococcal polysaccharide intercellular adhesin( PIA) • PIA – poly-N-acetyl glucosamine (PNAG) polymer • PIA polymers present in E. coli • The pga gene is involved with the formation of biofilms • This gene is similar to the staphylococcal gene ica • Now Yersinia pestis has been shown to make a gene homologous to this
Bap Protein • The biofilm associated protein • Structurally similiar to the surface proteins Esp of Enterococcus faecalis mus20 of Pseudomonas aeruginosa sty2875 of Salmonella typhi
Pseudomonas aeruginosa • Pseudomonas strains were thought to produce alginate • However alg mutants did not produce changes in the biofilm formation • Two new polymers have been found in Pseudomonas polymers • pelA-G gene produces a glucose rich polymer • pslA-O genes produce a mannose rich polymer
Population density and polysaccharide production • The connection between quorum sensing and biofilm architecture • Biofilm thickness seems to affect communication • Quorum sensing mechanism is not clearly defined but seems to be essential in the formation of the film and its channels
Vibrio cholera Major extracellular polysaccharide in the biofilm are VPS and the genes, vps VPS is negatively regulated by hapR. Hap R mutants produce rugose colonies and narrow channels in the biofilm architecture hapR gene encodes a transcription factor hapR is repressed by LuxO
Conjugative pili • Conjugative pili greatly accelerates initial adhesion and biofilm development by E. coli • Gram negative bacteria have adhesins at the tip of its fimbriae • E. coli responds to levels of nutrients and osmolarity
Adhesins • Adhesins are molecules that are attached to bacterial fimbriae
Bacterial Cell Characteristics • The phenotypes of the cells include: • a slower growth rate • increased antibiotic resistance • elevated frequency of lateral gene transfer
Quorum sensing • Is controlled by at least two different quorum sensing signals • Acyl-homoserine lactone CAI-1( cholera autoinducer 1) appears to play a significant role in biofilm formation • Under low cell density CAI levels are lose enough to permit LuxO mediated repression of hapR resulting in VPS production • This bacterium appears to initiate production of an extracellular matrix under conditions of low population density presumably before the establishment of a multicellular community
Antibiotic Resistance • In the center of the biofilm the bacteria have greater antibiotic resistance • Opsonisation of antibiotics • To resistance to phagocytosis
E. faecalis • E. faecalis biofilms on dental root canals, urethral catheters, uretheral stents ,and heart valves have been observed. • While it is not clear that the ability of E. faecalis to form biofilms is essential for virulence, it appears that a majority of clinical isolates do possess the ability to form a biofilm in vitro
Dental plaque • Genome – genome interactions:bacterial communities in initial dental plaque – Paul Kolenbrander et al – Trends in Microbiology. January 2005. • Found on the enamel of teeth • Epithelial cells of the oral mucosa • Participate in coaggregation which occurs between different species of bacteria
Plaque smear 400 different bacterial species can be found in plaque 1010bacteria/mg High levels of Ca++ and P Matrix forms with cells Dental Plaque - colonization
Supra gingival – above gums Subgingival –below the gums Also related to the tooth surface Definition
Primary Colonizers • The pellicle-coated tooth surface is colonized by Gram-positive bacteria such as Streptococcus sanguis, Streptococcus mutans, and Actinomyces viscosus • These organisms are examples of the "primary colonizers" of dental plaque. Bacterial surface molecules interact with components of the dental pellicle to enable the bacteria to attach or adhere to the pellicle-coated tooth surface.
Primary colonization • For example, specific protein molecules found as part of the bacterial fimbria (hair-like protein extensions from the bacterial cell surface) on both Streptococcus sanguis and Actinomyces viscosus interact with specific proteins of the pellicle (the proline-rich proteins) with a "lock and key" mechanism • This results in the bacteria firmly sticking to the pellicle-coating on the tooth surface (Mergenhagen et al. 1987). Within a short time after cleaning a tooth, these Gram-positive species may be found on the tooth surface.
Mechanisms for plaque formation • two distinct mechanisms: 1 ) the multiplication of bacteria already attached to the tooth surface, and 2) the subsequent attachment and multiplication of new bacterial species to cells of bacteria already present in the plaque mass.
Mechanisms • Two distinct mechanisms: • 1 ) the multiplication of bacteria already attached to the tooth surface • 2) the subsequent attachment and multiplication of new bacterial species to cells of bacteria already present in the plaque mass.
Complexity increases • The secondary colonizers include Gram-negative species such as Fusobacterium nucleatum, Prevotella intermedia, and Capnocytophaga species.
Tertiary colonizers • After one week of plaque accumulation, other Gram-negative species may also be present in plaque. These species represent what is considered to be the "tertiary colonizers", and include Porphyromonas gingivalis, Campylobacter rectus, Eikenella corrodens, Actinobacillus actinomycetemcomitans, and the oral spirochetes (Treponema species).
Characteristics • The structural characteristics of dental plaque in this time period reveal complex patterns of bacterial cells of cocci, rods, fusiform, filaments, and spirochetes. • In particular, specific associations of different bacterial forms have been observed. For example, the adherence of cocci to filaments results in a typical form referred to as "test-tube brushes" or "corn-cob" arrays • these structures can be seen in The structural interactions of the bacteria probably are a reflection of the complex metabolic interactions that are known to occur between different plaque microorganisms.
Coaggregation • Express components that mediate cell to cell binding • One cell type in a coaggregation partnerships, one cell type expresses a heat-inactivated protease sensitive surface adhesion • The other partner expresses a complementary heat stable protein
Biofilm Biochemistry • The production of succinic acid from Campylobacter species that is known to be used as a growth factor by Porphyromonas gingivalis. Streptococcus and Actinomyces species produce formate, which may then be used by Campylobacter species. • Fusobacterium species produce both thiamine and isobutyrate that may be used by spirochetes to support their growth. The metabolic and structural interactions between different plaque microorganisms are a reflection of the incredible complexity of this ecological niche.