Looking for the Best QSAR and Docking Methods
230 likes | 460 Vues
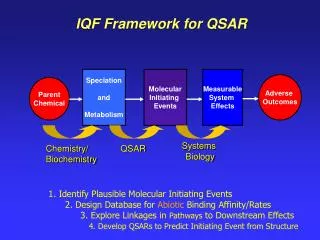
Looking for the Best QSAR and Docking Methods. Guillermo Restrepo Laboratorio de Química Teórica , Universidad de Pamplona, Pamplona, Colombia. Outline. Ranking How we rank Ranking problems QSAR models Docking programs Conclusions Acknowledgements. “Good”. “Bad”. 1. 2. 3. 4. 5.

Looking for the Best QSAR and Docking Methods
E N D
Presentation Transcript
Looking for the Best QSAR and Docking Methods Guillermo Restrepo Laboratorio de QuímicaTeórica, Universidad de Pamplona, Pamplona, Colombia
Outline • Ranking • How we rank • Ranking problems • QSAR models • Docking programs • Conclusions • Acknowledgements
“Good” “Bad”
1 2 3 4 5 6 We love rankings! La romería de San Isidro, Goya
How do we rank? • Priorities • • Subjectivities
Comparable x≥ y if all qi(x) > qi(y) or at least one attribute (qj) is higher for xwhile all others are equal. Incomparable If at least one qjfulfills qj(x) < qj(y) while the others are opposite (qi(x) ≥ qi (y)), x and y are incomparable. A B C D E F Hasse diagram Total set of linear extensions Brüggemann, R.; Restrepo, G.; Voigt, K. J. Chem. Inf. Model.2006, 46, 894-902.
rmn: ocurrence of object n at rank m A B C D E F 5 b d 4 3 e a 2 Average rank of n Av rkn= ∑mm∙pmn Ranking probability of having n at m c 1 pmn= rmn/ |LE| Restrepo, G.; Brüggemann, R.; Weckert, M.; Gerstmann, S.; Frank, H. MATCH Commun. Math. Comput. Chem.2008, 59, 555-584.
Best QSAR methods • Case study: • Mutagenicity • 95 aromatic & heteroaromatic amines • 13 QSAR models • Two statistics
Maran 1999 & Karelson 2000a are better than 10 other models. • It is not possible to state whether Karelson 2000b is better or worse than other models. • There are better models than Cash 2001 & Toropov 2001. Maran1999 Karelson2000a Karelson 2000b Basak 2001b Basak 1997 Vračko2004a,b Basak 2001a Cash 2005b Basak 1998 Cash 2001 Cash 2005a 594 linear extensions Toropov 2001 Restrepo, G.; Basak, S. C.; Mills, D. Curr. Comput-Aid Drug. 2011, 7, 109-121.
11 Maran1999 Karelson2000a 10 Basak 2001b 9 Basak 1997 8 Vračko2004a,b 7 Basak 2001a 6 Karelson 2000b Basak 1998 5 • Maran1999 & Karelson2000a and Basak 2001b are the less variable models. • Karelson 2000b & Cash 2005b are the most variable models. 4 Cash 2005b Cash 2005a 3 Cash 2001 2 Toropov 2001 1
Best Docking methods • Case study: • 10 docking programs: Dock4, DockIt, FlexX, Flo, Fred, Glide, Gold, LigFit, MOE, MVP • 8 protein targets • Two main characteristics: • prediction of conformations of small molecules bound to protein targets • virtual screening of compound databases to identify leads for a protein target Warren, G. L.; Andrews, C. W.; Capelli, A-M.; Clarke, B.; LaLonde, J.; Lambert, M. H.; Lindvall, M.; Nevins, N.; Semus, S. F.; Senger, S.; Tedesco, G.; Wall, I. D.; Woolven, J. M.; Peishoff, C. E.; Head, M. S. J. Med. Chem. 2006, 49, 5912-5931.
Protein-ligand conformations Percentage of compounds for which a docked pose was found within 2 Å of the crystal structure Nuclear hormone receptor Polypeptide deformilase Serine protease Polymerase Synthetase Isomerase Kinase 136 protein/ligand conformations
Protein-ligand conformations • There are better programs than MOE • There is no program behaving better than the others • Gold performs better than 4 other programs MVP Fred FlexX Glide Flo+ Gold Dock4 DockIt LigFit MOE 12,960 linear extensions
10 9 Gold 8 FlexX, Flo+, Glide 7 6 Fred, MVP 5 • All programs have variable positions in the ranking, except MOE. • The most suitable docking program to estimate protein-ligand conformations is Gold. 4 Dock4, DockIt, LigFit 3 2 MOE 1
Docking as a virtual screening tool Enrichment factor for actives (≤1 μM) found at 10% of the docking-score-ordered list Nuclear hormone receptor Metalloprotease Serine protease Metalloprotease Polymerase Synthetase Isomerase kinase
Docking as a virtual screening tool Ability to correctly identify all active chemotypes from a population of decoy molecules • MVP works better than 6 of the other programs • DockIt behaves worse than Flo+ and MVP • There is no program behaving better than all the others • Flex, Glide and Gold are the programs for which it is not possible to find a better or worse program Glide Gold FlexX MVP Dock4 Flo+ Fred LigFit MOE DockIt 259,200 linear extensions
10 MVP 9 8 7 Flo+ 6 FlexX, Glide, Gold 5 Dock4, Fred, LigFit, MOE 4 • All programs have quite variable positions in the ranking • The most suitable docking program to identify active chemotypes is MVP DockIt 3 2 1
Conclusions • With 2 statistics characterising QSAR models, we found 2 best models. • … and 2 “worse” models. • The docking program for protein-ligand conformations with the highest probability of being the best one (21%) is Gold. • MVP has 70% probability of being the best docking program for virtual screening searches.
Outlook • Why not using more statistics for QSAR models? • Instead of ordering Alice and Bob’s models, a work to do is to order QSAR models, e.g. linear & non-linear ones. • Some other attributes of QSAR methods need to be introduced, e.g. related to the applicability domain. • Computational costs and other docking programs features may be included in the study.
Acknowledgements Rainer Brüggemann Subhash C. Basak