DNA replication
DNA replication. Andy Howard Introductory Biochemistry 4 December 2008. DNA replication: accuracy!.


DNA replication
E N D
Presentation Transcript
DNA replication Andy HowardIntroductory Biochemistry4 December 2008
DNA replication: accuracy! • The extraordinary fidelity of heritance in prokaryotes and eukaryotes derives from the net accuracy of DNA replication. We’ll outline the steps of replication and the proofreading that goes with it. DNA replication
Prokaryotic DNA replication Semiconservative replication Unwinding of parent DNA Leading-strand replication Lagging-strand replication Okazaki fragments Eukaryotic replication Rates Multiple starting points Enzymes Other Repair & recombination Forms of repair Recombination Repair & Disease What we’ll discuss DNA replication
iClicker quiz • 1. Which of these DNA sequences is palindromic? • (a) TCGATG • (b) TAGGAT • (c ) GGCCCGGG • (d) TACGCGTA • (e) None of the above DNA replication
iClicker quiz #2 • Suppose someone introduced a drug that interfered with deacylation of H1. What effect would this have on chromatin assembly? • (a) It would prevent formation of the nucleosome core particle • (b) It would interfere with assembly of the structures connecting one core particle to the next • (c) Both (a) and (b) • (d) It would have no effect on chromatin assembly • (e) Like (c), except only in prokaryotes DNA replication
Semi-conservative replication Photo courtesy U. Costa Rica • A bit of a fanciful term • Refers to the fact that, during DNA replication, each daughter molecule contains one of the strands of the parent • Thus each daughter contains half (semi) of the original molecule • This mode of inheritance was predicted by the Watson/Crick model DNA replication
3 models (fig. 28.3) DNA replication
Meselson & Stahl • 1958: showed that DNA really is replicated this way • DNA grown with 15N has higher density • 15N DNA allowed to replicate exactly once has intermediate density DNA replication
Meselson-Stahl experiment • Fig. 28.4;note that the bottom 2 density gradients are for mixtures of generations DNA replication
The E.coli chromosome • One circular, double-stranded DNA molecule of about 4.6*106 bp • Replication begins in only one place, i.e. a single origin of replication (OriC in E.Coli) • Replication moves both directions until the two replication efforts meet at the termination site • Protein machine that accomplishes replication is the replisome;one replisome in each direction • Replication forks move 1000 bp/sec; thus E.coli can be replicated in 38 min = 2280 s DNA replication
By contrast: eukaryotes • Bigger chromosomes, more of them • Not circular, usually • Fruit-fly chromosomes: • Sex chromosome, 2 long autosomes,one tiny autosome • 1.65 * 108 bp, 14000 genes • Human • 22 pairs of autosomes, sex chromosome • 3.4*109 bp, ~22000 genes • Replication is bidirectional as in E.coli • More than one origin so replication is comparably fast even though rate is lower Drosophila chromosomeTEM reconstruction DNA replication
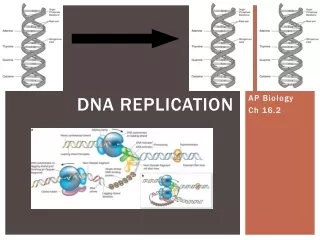
How replication works in prokaryotes • Takes place in the cytosol since there is no nucleus • Specific enzymes form the molecular machine to carry out the task • Has to involve separation of the strands • Process divided into initiation, elongation, and termination • Enzymatic functions identified for each segment DNA replication
Prokaryotic DNA polymerases • Several varieties • DNA polymerase III is the one responsible for most of the work (but the 3rd discovered);it’s the biggest and most complex • DNA pol I involved in error correction and helps with replication of one of the strands • DNA pol II also does DNA repair • Multi-subunit, complex entities DNA replication
Diagram from Kelman et al(1998) EMBO J. 17:2436 DNA Pol III DNA replication
Components of DNA Pol III DNA replication
So how does it work? • Add 1 nucleotide @ a time to 3’ end of growing chain • Substrate is a dNTP • Watson-Crick bp determines specificity • Enzyme spends 75% of time tossing out wrong bases • Forms phosphodiester linkage • Pol III remains bound to the replication fork Diagram from answers.com DNA replication
Error correction in DNA pol III • 3’-5’ proofreading recognizes incorrectly paired bases and repairs most of them • This is an exonuclease activity because it clips off the last nucleotide in the chain • 10-5 inherent error rate drops to 10-7 because the exonuclease goofs 1% of the time • Separate repair enzymes drop that down to 10-9 DNA replication
Processivity • This term refers to the fact that many nucleotides can be added to a growing chain following a single association event in which the polymerase (e.g. E.coli Pol III) associates with the template DNA. • We describe replication as highly processive if 50,000 bases can be replicated based on a single association of Pol III with our template. • subunits slide along, which is how this is done • complex is responsible for keeping the polymerase attached so that this is possible DNA replication
Initiation • Begins in E.coli at a single origin called OriC • DnaA binds to origin—region called DnaA box • Replication fork forms after it binds • Helicases & primasesset up for starting replication • Complementary RNA tag attached at the replication fork DNA replication
Elongation • DNA polymerase operates in 5’-3’ directionon both strands • For one strand that’s straightforward • replication moves in direction of unwinding of the DNA • This is known as leading-strand synthesis • For the other strand it must work opposite to the unwinding • therefore it’s more complicated • This is lagging-strand synthesis DNA replication
Termination • Replication needs to know how to stop • Defined sequence ter is opposite the origin on the chromosome • Specific enzyme, Tus, involved in recognizing termination signals • Ter has sequences that play a role in separating the daughter chromosomes Tus-Ter complex;images courtesy Memorial Univ., Newfoundland DNA replication
Leading-strand synthesis • One base at a time is incorporated by a subunit of DNA polymerase, complementary to existing strand • At some point RNA primer is replaced with DNA Image courtesy U.Pittsburgh DNA replication
iClicker question #3 3. A deleterious mutation in the gene coding for the subunit of DNA pol III would have what direct effect on replication in E.coli? • (a) it would increase error rates • (b) it would decrease processivity • (c ) it would prevent formation of Okazaki fragments • (d) none of the above DNA replication
Lagging-strand synthesis • Movement of enzyme is opposite to unwinding • Therefore it must work a few bases (~1000) at a time and then back up • The segments thus formed on the lagging strand are known as Okazaki fragments • DNA Pol I removes RNA primer • DNA ligases link together Okazaki fragments DNA replication
Primases • These are DNA-dependent RNA polymerase enzymes that initiate DNA synthesis, particularly on the lagging strand, where you need to do that at the beginning of each Okazaki fragment DnaG helicase binding domain PDB 2R6A347 kDa trimer ofheterotrimersBacillus stearothermophilus DNA replication
Role of DNA Polymerase I • Part of the system for producing a continuous DNA strand on the lagging side • Contains both 5’3’ polymerase activity and 3’5’ proofreading exonuclease activity • Also has 5’3’ exonuclease activity: that’s used to remove the RNA primer DNA replication
Rates and sequencing • Because there’s only one place where replication can begin, the process has to occur in discrete steps • The enzymes themselves are efficient, because they move with the unwinding of the double helix • Typical rates 1000 nucleotides/ sec • So for E.coli it takes 38 min = 2280 sec to replicate the entire chromosome(4.6*106 bp) / [(103 bp/sec)(2 directions)]=2300 sec DNA replication
Where are things happening? • Both leading- and lagging-strand synthesis are catalyzed in both the clockwise and counterclockwise directions. • Each DNA Pol III molecule is catalyzing both leading- and lagging-strand synthesis. DNA replication
Eukaryotic DNA polymerases image courtesy Memorial Univ., Newfoundland • More complex, as you’d expect • More than one initiation point • Therefore even though the enzymes work less rapidly, they can multiplex the process • The result is that human DNA can be replicated in roughly the same time scale as E.coli DNA DNA replication
Eukaryotic DNA polymerases • Several complexes involved: • a: Elongation and repair • d: Elongation and repair • e: Elongation and repair • b: DNA repair • g: Helps in replicating mitochondrial DNA • chloroplast polymerase • Don’t fall into the trap: these Greek letters don’t necessarily mean the same thing that they mean in the context of bacterial replication! Human DNA polymerase b: From answers.com DNA replication
Specific roles for a and d • d complex does leading-strand synthesis and 3’-5’ exonuclease activity, which is impressively effective • a and d involved in lagging strand synthesis: • a is DNA polymerase and RNA primase;d extends segment to complete Okazaki fragment DNA replication
Roles for DNA polymerase e Cryo EM image from Asturias et al (2006) Nature Struct Mol Biol13:35;yeast • Big,multi-subunit protein • Similar to DNA pol I from E.coli • It repairs and it fills gaps between Okazaki fragments • Largest piece is a polymeraseand does 3’5’ proofreading DNA replication
iClicker question 4 4. A deleterious mutation in eukaryotic DNA polymerase would directly affect • (a) trafficking of membrane-bound proteins from the ribosome to the plasma membrane • (b) assembly of the nucleosome core particle • (c) energy production in the cell • (d) harvesting of light from the sun • (e) there is no eukaryotic DNA polymerase DNA replication
DNA repair • DNA is the only macromolecule that gets repaired: it’s too important not to • A single base error can be fatal, even in prokaryotes • Natural rates of misincorporation are small but nonzero • Rate can go up upon exposure to ionizing radiation, some chemicals, some toxins DNA replication
Direct repair • Enzymes scan DNA for particular lesions • Pyrimidine dimers are noted and repaired this way • Some can replace the base without breaking the phosphodiester backbone Image courtesy U. München DNA replication
Excision repair • Endonuclease recognizes lesion • Cleaves upstream &downstream —12-13 bases • Only cleaves damaged strand • Removal may require helicase • DNA polymerase (I in prokaryotes) fills the gap • DNA ligase reseals the lesion Diagram courtesy Beth Montelone, Kansas State U. DNA replication
Other repairs H2O • Repairing hydrolytic deamination of A, C, G: • DNA glycosylase flips base out and hydrolyzes glycosidic bond • Endonuclease sutures in one replacement base (sometimes part of same protein) NH3 DNA replication
Recombination • Recombination is any exchange or transfer of DNA from one spot to another • Homologous recombination involves exchanges in closely-related sequences; can involve paired chromosomes • Transposons are elements that can be readily recombined nonhomologously DNA replication
Holliday model • Cf.fig.28.19 • Involves nicking, then rotation • Can result in exchanging the ends of two homologous chromosomes DNA replication
Recombination in E.coli • RecBCD endonuclease creates single-stranded DNA with free 3’ • SS DNA invades double helix of neighboring DNA • RecA promotes strand exchange: • Requires energy • Formation of triple-stranded intermediate • Branch migrates down chain Mazin& Kowalczykowski,(1998) EMBO J.17:1161 DNA replication
Recombination as repair • Bad lesions are simply skipped • Intact strand from one daughter acts as template for repairing broken strand • Most recombination genes play roles in repair too E.coli RecA in compressed helical form DNA replication
Repair & Disease • Repair deficiencies render the organism susceptible to mutation-related maladies • BRCA1&BRCA2 are proteins that bind to RecA in humans and help repair DSBs • Therefore mutations in these proteins leave people prone to cancer PDB 1t15:BRCA1 BRCT domains complexed to BACH1 helicase DNA replication
PDB 2csv Ataxia telangiectasia • Ataxia is lack of coordination • Telangiectasia is spider-veins • Condition involves both symptoms • Caused by mutation in specific protein involved in cell-cycle control • Patients are abnormally prone to cancer, other DNA-repair related conditions • Ataxia telangiectasia involves failure to recognize DSBs • ~1% population heterozygous • 8-10% of breast cancer patients are heterozygotes for this condition; Swift (2001) JNCI93:84 DNA replication