miRDeep2*
Supplementary Figure 2 : . 2. genomic reads. smallRNA reads. 1. blast. 3. matching genomic reads/mates. de novo assembly. 4. 5. prepare miRNA-candidates reference. miRDeep2*. 6. miRNA complement.

miRDeep2*
E N D
Presentation Transcript
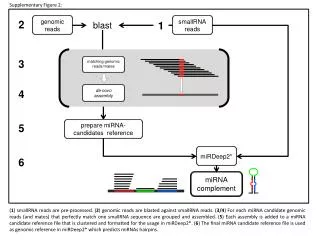
Supplementary Figure 2: 2 genomic reads smallRNA reads 1 blast 3 matching genomic reads/mates de novo assembly 4 5 prepare miRNA-candidates reference miRDeep2* 6 miRNA complement (1) smallRNA reads are pre-processed. (2) genomic reads are blasted against smallRNA reads. (3/4) For each miRNA candidate genomic reads (and mates) that perfectly match one smallRNA sequence are grouped and assembled. (5) Each assembly is added to a miRNA candidate reference file that is clustered and formatted for the usage in miRDeep2*. (6) The final miRNA candidate reference file is used as genomic reference in miRDeep2* which predicts miRNAs hairpins.
Supplementary Figure 3: genomic reads smallRNA reads blast miRNA components from one hairpin hit different sets of genomic reads 1 2 3 independent de novo assembly for each group of genomic reads Different contigs that are divergent in length but contain same hairpin contig 3 contig 2 contig 1 MiRNA hairpins contribute with at least three kinds of reads to the blast search: mature-, seed- and star-sequence-reads. The correponding genomic matches will be assembled in divergent contigs by MirCandRef. They represent different parts of the same locus, each with the identical miRNA-hairpin sequence. This leads to a high level of redundancy in the MiRCandRef contigs that is handled by clustering of the contigs and modifications to miRDeep2.