What is the NMR Assignment Issue?
940 likes | 1.33k Vues
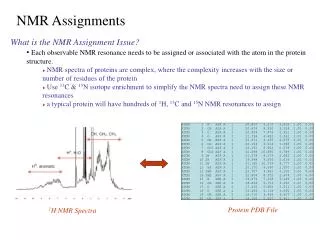
NMR Assignments. What is the NMR Assignment Issue? Each observable NMR resonance needs to be assigned or associated with the atom in the protein structure. NMR spectra of proteins are complex, where the complexity increases with the size or number of residues of the protein

What is the NMR Assignment Issue?
E N D
Presentation Transcript
NMR Assignments • What is the NMR Assignment Issue? • Each observable NMR resonance needs to be assigned or associated with the atom in the protein structure. • NMR spectra of proteins are complex, where the complexity increases with the size or number of residues of the protein • Use 13C & 15N isotope enrichment to simplify the NMR spectra need to assign these NMR resonances • a typical protein will have hundreds of 1H, 13C and 15N NMR resonances to assign Protein PDB File 1H NMR Spectra
NMR Assignments Again, as illustrated here, the goal is to explicitly assign each H, C, & N in the protein’s primary sequence with its corresponding NMR resonance 15N 114.8 ppm HN 7.08 ppm 15N 125.6 ppm HN 8.20 ppm 13Ca 58.6 ppm Ha 4.09 ppm 15N 119.3 ppm HN 7.76 ppm 13Ca 55.5 ppm Ha 3.76 ppm 13CO171.9 ppm 13CO178.1 ppm 13CO170.9 ppm 13Cb 17.5 ppm Hb 1.45 ppm 13Cb 42.9 ppm Hb 1.52 ppm 13Cb 64.8 ppm Hb 3.73 ppm 13Cg 27.9 ppm Hg 1.65 ppm 13Ca 59.9 ppm Ha 4.35 ppm 13Cd 25.4 ppm; 25.7 ppm Hd 0.82 ppm; 0.98 ppm
NMR Assignments • Predicting NMR Chemical Shifts • A ever-growing number of computer programs are being developed to predict chemical shifts from structure or sequence. • SHIFTS, SHIFTX2, SPARTA+, Camshift, PPM, 4DSPOT, shAIC, etc. • Empirical models based on high quality structures with NMR assignments, and molecular dynamics J. Biomol. NMR 2010 48(1):13. J. Biomol. NMR 2012 54(3):257
NMR Assignments • How Are NMR Assignments Made For a Protein? • Requires the collection and analysis of multidimensional NMR data • 2D, 3D, 4D NMR spectra • This in turns requires software to assist in the processing and analysis of the data • ongoing effort to develop software to automate NMR assignments • not “100%” efficient but significantly aids in the manual assignment Assignment Table . . .
NMR Assignments • NMR Data Processing Software • Needs to specifically handle format of multidimensional NMR data • 2D, 3D, 4D NMR spectra • NMRPipe, Felix, ACD and others • all have similar functions and capability • all handle common instrument data formats (Bruker, Varian) • choice is primarily based on personal preference • NMRpipe: • UNIX/LINUX • simple script to process NMR data • mimics flow of processing steps • uses UNIX pipe functionality to pass data between one function to the next
NMR Assignments • NMR Data Processing Software • Main steps in the processing process include: • window function (SP), zero fill (ZF), Fourier transform (FT), phase (PS), transpose (TP) • Other steps include • removing solvent (SOL), linear prediction (LP) and data extraction (EXT) • These steps are simply repeated for each dimension of the • NMR data Standard Processing Script for 3D NMR Data X Processing steps for X,Y,Z dimensions of 3D spectra Y Z
NMR Assignments • NMR Data Processing Software • Because of the exponential increase in time to collect nD NMR spectra, the number of data points collected for the indirect FIDs are kept to a minimum • 1D NMR ~few mins. 2D ~few hours 3D ~ few days • 1D NMR 8-32K pts 2D 2K x 512 pts 3D 2K x 128 x 80 pts • Two major impacts: • FIDs in indirect dimension are typically truncated artifacts in the spectra • FIDs in indirect dimension have very low resolution • These issues are addressed in processing the data • ZF, SP, LP FT
NMR Assignments • NMR Data Processing Software • A main goal in applying a window function for a nD NMR spectra is to remove the truncation by forcing the FID to zero. Truncated FID with spectra “wiggles” Apodized FID removes truncation and wiggles
NMR Assignments • NMR Data Processing Software • Some common window functions with the corresponding NMRPipe command
NMR Assignments • NMR Data Processing Software • Want to maximize digital resolution, number of data points in each dimension • time constraints are a practical limitation for nD NMR data
NMR Assignments • NMR Data Processing Software • Improve digital resolution by adding zero data points at end of FID • essential for nD NMR data • no significant gain after one ZF, just interpolation between points 8K data 8K zero-fill 8K FID 16K FID 8K zero-filling No zero-filling
NMR Assignments • NMR Data Processing Software • Linear Prediction • extrapolate FID data in time domain • enhances resolution • works best for data without significant relaxation • assumes sinusoid shape • a set of coefficients is found such that linear combination of a group of points predicts the next point in the series. • number of coefficients determine the number of NMR signals (damped sinusoids) that can be predicted • LP is usually limited to extending data to about twice its original size • forwardlinearprediction - points immediately after each group are predicted • backwardlinearprediction - points immediately before each group are predicted • forward-backwardlinearprediction - combines results from separate forward- and backward-linear prediction calculations. LP
NMR Assignments • NMR Data Processing Software • Linear Prediction • model (set of coefficient) can be applied to predict a new synthetic point • uses a group of existing points from the original data • new point along with group from the original data is used to predict yet another point • process can be continued indefinitely • becomes unstable when group contains all synthetic points • Mirror Image LP • LP order (number of coefficients) must be as large as the number of signals to extract, but smaller than half the original data size. • For constant time data, (no decay) can temporarily add the data's mirror image complex conjugate for the LP calculation and then discard it. • time increment must be the same between each point • either 0,0 or 90,-180 phase correction LP Progress in Nuclear Magnetic Resonance Spectroscopy (1988), 20(6),515-626
NMR Assignments • NMR Data Processing Software • Effects of Combining Linear Prediction with Zero Filling • significant improvement in resolution for nD NMR data collected with minimal data points
NMR Assignments • NMR Data Processing Software • uniform data sampling • avoids under-sampling frequencies • FT algorithms expect uniform spacing of digital data The Nyquist theorem Need to sample twice as fast (DW)as the fastest frequency Traditional NMR acquires EVERY data point with a uniform time-step between points. Reason why nD NMR experiments take so long, why FIDs in indirect dimensions are truncated and the spectra have low resolution and sensitivity
NMR Assignments • NMR Data Processing Software • Non-uniform data sampling • significant improvement in resolution and sensitivity for nD NMR data • Don’t need uniform sampling, just need alternative to FFT to process the data. • The sampling non-uniform scheme is the primary decision and impact on the spectra Exponential in both t1 and t2 exponential in t1 and linear in t2 randomly sampled from an exponential distribution in t1 and t2 Random in t1 and t2. Graham A. Webb (ed.), Modern Magnetic Resonance, 1305–1311.
NMR Assignments • NMR Data Processing Software • Non-uniform data sampling • VERY IMPORTANT POINT, tn is no longer defined by DW and number of points • tnis now user defined since DW is no longer relevant. • Avoid FID truncation, maximize resolution voltage time Traditional NMR FID is truncated because number of points and DW determine how much of the FID can be collected NUS NMR FID is under-sampled, but the entire FID is sampled.
NMR Assignments • NMR Data Processing Software • Non-uniform data sampling • Both noise (N) and signal to noise (SNR) are proportional to the total evolution time • Optimal setting is 1.3T2 of the evolving coherence • Maximize sensitivity Magn. Reson. Chem. 2011, 49, 483–491
NMR Assignments • NMR Data Processing Software • Non-uniform data sampling • What is the optimal sampling density? • Increase enhancement by increase exponential bias, eventually regenerate truncated FID • Highly resolved spectra is pT2 • TSMP – time constant for the exponential • weighting of the sampling. • - enhancement • lw – line width Magn. Reson. Chem. 2011, 49, 483–491
NMR Assignments • NMR Data Processing Software • Non-uniform data sampling • A 1.5 to 2.0 bias to early data points and a 4x reduction yields a 2x enhancement • Or a 3T2 with a 3x reduction yields a 1.7 enhancement Truncated FID Sampling Density/LW = TSMP/T2 Magn. Reson. Chem. 2011, 49, 483–491
NMR Assignments • NMR Data Processing Software • Non-uniform data sampling • Different sampling schemes have different performances at different sampling densities • Sinusoidal Poisson Gap is currently the best – random sampling, while minimizing gap size particularly at the beginning and end of the FID • Some drastic sampling densities at 1% or less. Top Curr Chem. 2012 ; 316: 125–148
NMR Assignments • NMR Data Processing Software • Non-uniform data sampling • Dramatic gain in resolution for 48 kDa protein with only 3% sampling of the Nyquist matrix • Same experimental time for US and NUS J Biomol NMR. 2009 November; 45(3): 283–294.
NMR Data Processing Software • Non-uniform data sampling • How is the time-domain data processed? • Use the partial data to reconstruct the full Nyquist grid then process as normal (nmrPipe) • maximum entropy reconstruction is a common approach • forward maximum entropy (FM), fast maximum likelihood reconstruction (FMLR) • multi-dimensional decomposition (MDD); and compressed sensing (CS) • MddNMR:http://www.enmr.eu/webportal/mdd.html • Newton:http://newton.nmrfam.wisc.edu/newton/static_web/index.html • RNMRTK:http://rnmrtk.uchc.edu/rnmrtk/RNMRTK.html • mpiPipe: Available by contacting the Wagner Group
NMR Assignments • NMR Data Processing Software • Solvent Removal (SOL) • protein NMR spectra are typical collected in water • the large solvent signal can interfere with the interpretation of the NMR data • Carrier frequency is usually centered on the water signal • the signal associated with the water resonance can be filtered or subtracted from the time domain of the FID SOL
NMR Assignments • NMR Data Processing Software • Solvent Removal (SOL) without Solvent Subtraction with Solvent Subtraction
NMR Assignments • NMR Data Processing Software • Phase Correction (PS) • Because of the challenges of phasing nD NMR data and the baseline artifacts that first-order phase corrections are known to cause, typically phase corrections are set to 0,0 or 90-180 by proper delays in the pulse sequence • A number of methods of data collection are used to obtain phase correction in the indirect dimensions • Fourier transformed data contains a real part that is an absorption lorentzian and an imaginary part which is a dispersion lorentzian • we want to maintain the real absorption mode line-shape • done by applying a phase factor (exp(iQ)) to set F to zero • this is what we are doing when we phase the spectra
NMR Assignments • NMR Data Processing Software • Phase Correction (PS) • Phase of the peak is determined by the relative phase of the pulse and the receiver • to obtain correct phasing in the indirect dimension, we need to collect both sine and cosine modulated data • alternate both the phase of the pulse relative to the receiver and the storage of this data between real (sine) and imaginary (cosine)
NMR Assignments • NMR Data Processing Software • Phase Correction (PS) • Phase of the peak is determined by the relative phase of the pulse and the receiver • Also determines the order in which the data is stored. • Some Common Phase Cycle Schemes: • STATES – phase cycles the 90o-pulses prior to t1 incrimination by 900 • TPPI – phase cycles both the receiver and the 90o-pulses prior to t1 by 90o for each t1 increment • States-TPPI – phase cycles both the receiver and the 90o-pulses prior to t1 by 180o for each t1 increment • Echo-antiecho – uses gradients to reduce the number of phase cycling steps and combines N (echo) and P(antiecho) coherence selection
NMR Assignments • NMR Data Processing Software • Phase Correction (PS)
NMR Assignments • NMR Data Processing Software • Phase Correction (PS) The phase introduced by a gradient of duration τG to coherence of order p which involves k spins with gyromagnetic ratios gk is given by: φ(r) = r Gz τG Sk( pkγk) Complex Fourier transformation and combination of the two signals yields a purely absorptive spectrum with frequency sign discrimination.
NMR Assignments • NMR Data Processing Software • Data Conversion (bruk2pipe) • Prior to processing the NMR data by NMRPipe is a requirement to convert the file format • This process requires defining some important experimental parameters • number of points, sweep width, phase cycling, etc Phase cycling determines how the data is stored and retrieved • States - odd data points are written to the real data array, even data points to the imaginary data array. source 1 2 3 4 = real 1 3 + imaginary 2 4 • TPPI - data are copied to the real data array. source 1 2 3 4 = real 1 2 3 4 • Echo-antiecho - 4 data points are mixed and written to the real and imaginary data arrays. source 1 2 3 4 = real 1+3 4-2 + imaginary 2+4 1-3 • States-TPPI - Same as States, but every second real and imaginary data point is multiplied by -1. source 1 2 3 4 = real 1 -3 + imaginary 2 -4
NMR Assignments • NMR Data Processing Software • NMR data analysis/visualization • NMRDraw, NMRViewJ, PIPP, etc • Again, most programs have similar functionality, choice is based on personal preference • display the data (zoom, traces, step through multiple spectra, etc) • Peak-picking – identify the X,Y or X,Y,Z or X,Y,Z,A chemical shift coordinate positions for each peak in the nD NMR spectra Peak Picking List
NMR Assignments • NMR Data Processing Software • NMR data analysis/visualization • Peak Picking • Critical for obtaining accurate NMR assignments • Especially for software for automated assignments • Only provide primary sequence and peak-pick tables • Two General Approaches to Peak Picking • Manual • time consuming • can evaluate crowded regions more effectively • Automated • pick peaks above noise threshold OR • pick peaks above threshold with characteristic peak shape • only about 70-80% efficient • crowded overlap regions and noise regions (solvent, T2 ridges) cause problems • noise peaks and missing real peaks cause problems in automated assignment software J. OF MAG. RES. 135, 288–297 (1998)
NMR Assignments • NMR Data Processing Software • NMR data analysis/visualization • What is the Statistical likelihood that a signal is a peak? 100 simulated spectra containing a single peak with random noise. A successful identification occurred if the known peak has the highest intensity that is at least 1.414 times greater than the next intense peak. A signal intensity of 1 corresponds to a SNR of 80. J Biomol NMR (2013) 55:167–178.
NMR Assignments • NMR Data Processing Software • Automated NMR assignments • AutoAssign, CONTRAST, GARANT, PASTA, etc • uses peak lists, primary protein sequence, details of NMR experiments • tries to mimic “skilled user”, uses databases of previous assignments, etc • Automated analysis of NOESY data is a sub-set of the NMR assignment issue with programs designed to specifically address this need • AutoStructure, CANDID, ARIA, ROSSETTA, etc From, peak-lists and protein sequence, software attempts to make the assignment. Not 100% success rate, still need user intervention to complete/correct assignments. Most problems arise from quality of peak-list: noise, missing peaks, etc. Need to Know How Assignments are Made!
NMR Assignments dbN dbN daN daN dNN dNN dNN • NMR Assignment Protocol • 2D NMR Experiments • Kurt WüthrichNobel prize in 2002for developing NMR to determine 3D structures of proteins. • Wüthrich “NMR of Proteins and Nucleic Acids” 1986, John Wiley & Sons • Applicable for proteins of <100 amino acids • Primarily dependent on three 2D experiments: NOESY, COSY, TOCSY • Sequence-Specific Resonance Assignments in Proteins (Backbone Assignemnts) Takes advantage of short sequential distances between CaiH, CbiH and NHi+1
NMR Assignments • 2D NMR Experiments • 2D COSY • Correlation Spectroscopy • Correlates 1H resonances that are scalar coupled (3J) • Identifies which NHi resonances are bonded to CaHi resonances • separated by three-bonds • chemical shift evolution based on J occurs during t1 • requires the sample be in H2O (90/10 H2O/D2O) to observe NH • all three-bond couplings observed, not just NH-Ca • spectra is symmetric • strength of cross peak depends on strength of coupling constants • all predicted peaks are not necessarily observed • weak couplings • obscured by solvent, noise • overlap or degenerate peaks
NMR Assignments • 2D NMR Experiments • 2D COSY • Typical Small Protein COSY
NMR Assignments • 2D NMR Experiments • 2D NOESY • Nuclear Overhauser Spectroscopy • Correlates 1H resonances that close in space (≤5Å) • also contains COSY peaks • NOE intensity builds up during mixing time (tm), ususally 100-150 ms • Correlates NHi+1 resonances with CaHi resonances
NMR Assignments • 2D NMR Experiments • 2D NOESY • Typical Protein NOESY (Lysozyme) Both NHi-Cai and NHi+1-Cai are present
NMR Assignments • 2D NMR Experiments • Making the Sequential Assignments • Connecting COSY (NHi-Cai) peaks with NOESY (NHi+1-Cai) • COSY experiment allows you to identify the NHi-Cai cross peaks in the NOESY experiment • N-terminal amino acid only has one cross peak associated with its NH chemical shift The Backbone Walk NOESY cross peak COSY cross peak NHi+1-Cai A24 NHi-Cai NHi-Cai NHi+1-Cai T27 Y28 NHi-Cai NHi+1-Cai F25 NHi-Cai D26 NHi+1-Cai NHi-Cai A24 F25 T27 Y28 D26 Biochemistry 1989, 28, 1048-1054 NH Chemical Shifts (ppm)
NMR Assignments • 2D NMR Experiments • Verifying the Sequential Assignments and Side-Chain Assignments • The accuracy of the backbone assignments from connecting COSY (NHi-Cai) peaks with NOESY (NHi+1-Cai) can be verified by proper assignment of the side-chain with the backbone assignments. • know the primary sequence of the protein • therefore, know what amino acid is residue (i) and what amino-acid should be (i+1) • amino acid type indicates the number and type or chemical shifts that should be observed for the residue As example: Gly – no side chain Ala – single methyl (1.39 ppm) Val – two g methlys (0.97 & 0.94 ppm) one Hb (2.13 ppm)
NMR Assignments • 2D NMR Experiments • Connectivity Patterns • COSY TOCSY patterns • for the 20 amino acids • Side-chain assignments • involves “matching” • the expected patterns • and typical chemical • shift ranges • Some connectivity • patterns are not unique • and can only eliminate • some possible • assignments In real data, overlapping or missing cross-peaks are common. Connectivity pattern may not exactly match predicted.
NMR Assignments • 2D NMR Experiments • Connectivity Patterns Leu - expected Cd Cg Cb Ca Leu - actual Cb/Cg Cb Cd Ca Structure induces chemical shift changes which perturbs the pattern and induces overlap. But, the data has to be consistent with the amino-acid spin system or the assignment is probably incorrect
NMR Assignments • 2D NMR Experiments • Connectivity Patterns • NMR assignments should be consistent with expected trends • significant differences should be explained by the structure • (ring current, h-bonds, etc)
NMR Assignments • 2D NMR Experiments • 2D TOCSY • TOtal Correlation SpectroscopY • cross peaks are generated between all members of a coupled spin network • NMR resonances for the complete side-chain spin systems is obtained • coherence transfer period occurs during a multi-pulse spin-lock period • length of spin-lock determines how “far” the spin coupling network will be probed • 1/(10 JHH) should be used for each transfer step • not all correlations are observed COSY TOCSY Spin-Lock Pulse (~14ms)
NMR Assignments • 2D NMR Experiments • 2D TOCSY • What happens during the spin-lock time cannot be described in terms of vector models or product • operators, because it relies on strong coupling • Under strong coupling, chemical shift differences between different spins become negligible • Two states ab and ba become identical in energy • Instead of transition of single spins, the coherences now involves transitions of combinations of spins • Under this condition, a coherence of one spin is actually in resonance with a coherence of its coupling partner(s) (all with the same frequency), and will oscillate back and forth between all coupled spins
NMR Assignments • 2D NMR Experiments • 2D TOCSY • Typical Small Protein TOCSY • Side-chain spin systems are correlated with NH resonance Boxed regions indicate side-chain spin systems for His and Ile, respectively Bull. Korean Chem. Soc. 2001, Vol. 22, No. 5 507
NMR Assignments • 3D NMR Experiments • Takes advantage of 13C and 15N labeling • Extends assignments to proteins in the 20-25 kDa range • Extends Connectivity by Scalar Coupling (J) into 3D dimensions • Primarily uses one-bond heteronuclear coupling (1H-13C, 1H-15N) • 1J generally stronger than 3J • 2D 1H-15N HSQC is the root experiment of most of the standard triple-resonance (1H, 13C, 15N) NMR experiments • 3D NMR simplifies data and removes overlap by spreading information into third dimension • Requires multiple experiments (≥ 6) to “walk through” the backbone assignments similar to the • 2D COSY & NOESY experiments • Requires a similar number of additional experiments to obtain the side-chain assignments
NMR Assignments • 3D NMR Experiments • 2D 1H-15N HSQC experiment • correlates backbone amide 15N through one-bond coupling to amide 1H • in principal, each amino acid in the protein sequence will exhibit one peak in the 1H-15N • HSQC spectra • also contains side-chain NH2s (ASN,GLN) and NeH (Trp) • position in HSQC depends on local structure and sequence • no peaks for proline (no NH) Side-chain NH2