Data Workflow Management, Data Preservation and Stewardship
640 likes | 676 Vues
Learn how scientific data workflows streamline processes, automate tasks, and optimize data analysis. Explore challenges and benefits, discover Kepler's role, and delve into distributed execution concepts.

Data Workflow Management, Data Preservation and Stewardship
E N D
Presentation Transcript
Data Workflow Management, Data Preservation and Stewardship Greg Hughes and Peter Fox Data Science – ITEC/CSCI/ERTH-4350/6350 Module 9, Week 11, November 15, 2016
Contents • Reading – none from last week • Scientific Data Workflows • Data Stewardship • Summary • Next class(es) • Projects
Scientific Data Workflows • What are they? • Why you would use them? • Some more detail in the context of Kepler • www.kepler-project.org • Some pointers to other workflow systems
What is a workflow? • General definition: series of tasks performed to produce a final outcome • E.g. following a recipe to bake a pie • Scientific workflow – “data analysis pipeline” • Automate tedious jobs that scientists traditionally performed by hand for each dataset • Process large volumes of data faster than scientists could do by hand
Background: Business Workflows • Example: planning a trip • Need to perform a series of tasks: book a flight, reserve a hotel room, arrange for a rental car, etc. • Each task may depend on outcome of previous task • Days you reserve the hotel depend on days of the flight • If hotel has shuttle service, may not need to rent a car • E.g. tripit.com
What about scientific workflows? • Perform a set of transformations/ operations on a scientific dataset • Examples (cf. Data Mining examples) • Generating images from raw data • Identifying areas of interest in a large dataset • Classifying set of objects • Querying a web service for more information on a set of objects • Many others…
More on Scientific Workflows • Formal models of the flow of data among processing components • May be simple and linear, or more complex • Can process many data types: • Archived data • Streaming sensor data • Images (e.g., medical or satellite) • Simulation output • Observational data
Challenges • Questions: • What may be some challenges for scientists implementing scientific workflows? • What are some challenges to executing these workflows?
Challenges • Mastering a programming language • Visualizing the workflow, end-to-end • Sharing/exchanging that workflow, updating it • Dealing with data formatting • Locating datasets, services, or functions (input and outputs, metadata…)
E.g. Kepler Scientific Workflow Management System • Graphical interface for developing and executing scientific workflows • Scientists create workflows by dragging and dropping • Automates low-level data processing tasks • Provides access to data repositories, compute resources, workflow libraries
Additional Benefits of Scientific Workflows • Documentation of aspects of analysis • Visual communication of analytical steps • Ease of testing/debugging • Reproducibility • Reuse of part or all of workflow in a different project
Additional Benefits • Integration of multiple computing (deployment) environments • Automated access to distributed resources via web services and Cloud technologies • System functionality to assist with integration of heterogeneous components
Why not just use a script? • Scripts do not (usually) specify low-level task scheduling and communication • Often are very platform-dependent • -> Can’t be easily reused? • May not have sufficient documentation to be adapted for another purpose
Why is a GUI useful? • No need to learn a programming language • Visual representation of what workflow does • Allows you to monitor workflow execution • Enables user interaction • Facilitates sharing of workflows
The Kepler Project • Goals • Produce an open-source scientific workflow system • enable scientists to design scientific workflows and execute them • Support scientists in a variety of disciplines • e.g., biology, ecology, astronomy • Important features • access to scientific data • flexible means for executing complex analyses • enable use of Grid-based approaches to distributed computation • semantic models of scientific tasks • effective UI for workflow design
Distributed execution • Opportunities for parallel execution • Fine-grained parallelism • Coarse-grained parallelism • Few or no cycles • Limited dependencies among components • ‘Trivially parallel’ • Many science problems fit this mold • parameter sweep, iteration of stochastic models • Current ‘plumbing’ approaches to distributed execution • workflow acts as a controller • stages data resources • writes job description files • controls execution of jobs on nodes • requires expert understanding of the Grid system • Scientists need to focus on just the computations • try to avoid plumbing as much as possible
Distributed Kepler • Higher-order component for executing a model on one or more remote nodes • Main and client controllers handle setup and communication among nodes, and establish data channels • Extremely easy for scientist to utilize • requires no knowledge of grid computing systems IN OUT Controller Controller Main Client
Managing Data Heterogeneity? • Data comes from heterogeneous sources • Real-world observations • Spatial-temporal contexts • Collection/measurement protocols and procedures • Many representations for thesame information (count, area, density) • Data, Syntax, Schema, Semantic heterogeneity • Discovery and “synthesis” (integration) performed manually • Discovery often based on intuitive notion of “what is out there” • Synthesis of data is very time consuming, and limits use
A simple Kepler workflow Composite Component (Sub-workflow) Loops often used in SWFs; e.g., in genomics and bioinformatics (collections of data, nested data, statistical regressions, ...) (T. McPhillips)
A simple Kepler workflow Lists Nexus filesto process (project) Reads text files Parses Nexus format Draws phylogenetic trees PhylipPars infers trees from discrete, multi-state characters. Workflow runs PhylipPars iteratively to discover all of the most parsimonious trees. UniqueTrees discards redundant trees in each collection. (T. McPhillips)
A simple Kepler workflow An example workflow run, executed as a Dataflow Process Network
Navigate errors and warnings within the workflow Search for and insert “adapters” to fix (structural and semantic) errors … Statically perform semantic and structural type checking Workflow validation in Kepler
+ Provenance Framework • Provenance • Tracks origin and derivation information about scientific workflows, their runs and derived information (datasets, metadata…) • Types of Provenance Information: • Data provenance • Intermediate and end results including files and db references • Process (=workflow instance) provenance • Keep the workflow definition with data and parameters used in the run • Error and execution logs • Workflow-design provenance
Kepler Provenance Recording Utility • Parametric and customizable • Different report formats • Variable levels of detail • Verbose-all, verbose-some, medium, on error • Multiple cache destinations • Saves information on • User name, Date, Run, etc…
Some other workflow systems • SCIRun • Sciflo • Triana • Taverna • Pegasus • Some commercial tools: • Windows Workflow Foundation • Mac OS X Automator • http://www.isi.edu/~gil/AAAI08TutorialSlides/5-Survey.pdf • See reading for this week
Workflows • Important for your last assignment….
Now …. Data Stewardship • Combining multiple data life cycle, management aspects together • Keep the ideas in mind as you complete your assignments • Why it is important • Some examples
Why it is important • Need ability to read the underlying sources, e.g. the data formats, metadata formats, knowledge formats, etc. • Need ability to know the inter-relations, assumptions and missing information • We’ll look at a (data) use case for this shortly • But first we will look at what, how and who in terms of the full life cycle
What to collect? • Documentation • Metadata • Provenance • Ancillary Information • Knowledge
Who does this? • Roles: • Data creator • Data analyst • Data manager • Data curator
How it is done • Opening and examining Archive Information Packages! Yes, people look at them. • Reviewing data management plans and documentation! Yes, people look at them. • Talking (!) to the people: • Data creator • Data analyst • Data manager • Data curator • Sometimes, reading the data and the code
Data-Information-Knowledge Ecosystem Producers Consumers Experience Data Information Knowledge Creation Gathering Presentation Organization Integration Conversation Context
Remember - Acquisition • Learn / read what you can about the developer of the means of acquisition • Documents may not be easy to find • Remember bias!!! • Document things as you go • Have a checklist (see the Data Management list) and review it often
Curation • Consider the “changes” in the organization and presentation of the data • Document what has been (and has not been) done • Consider and address the provenance of the data to date, you are now THE next person in the workflow… • Be as technology-neutral as possible • Look to add information and meta-information
Preservation • Usually refers to the latter part of data life cycle • Archiving is only one component • Intent is that ‘you can open it any time in the future’ and that ‘it will be there’ • This involves steps that may not be conventionally thought of • Think 10, 20, 50, 200 years (or 1 hour!) …. looking historically gives some guide to future considerations
Some examples and experience • NASA, NOAA • http://wiki.esipfed.org/index.php/Preservation_and_Stewardship • Library community • Note: • Mostly in relation to publications, books, etc. but some for data • Note that knowledge is in publications but the structural form is meant for humans not computers, despite advances in text analysis, NLP • Very little for the type of knowledge -> data we are considering: i.e. in machine accessible form
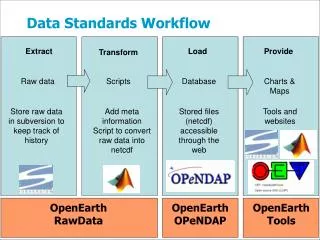
Use case: a real live one; deals mostly with structure and (some) content HDF4 Format "Maps"for Long Term Readability C. Lynnes, GES DISC R. Duerr and J. Crider, NSIDC M. Yang and P. Cao, The HDF Group HDF=Hierarchical Data Format NSIDC=National Snow and Ice Data Center GES=Goddard Earth Science DISC=Data and Information Service Center
In the year 2025... A user of HDF-4 data will run into the following likely hurdles: • The HDF-4 API and utilities are no longer supported... • ...now that we are at HDF-7 • The archived API binary does not work on today's OS's • ...like Android 9.1 • The source does not compile on the current OS • ...or is it the compiler version, gcc v. 12.x? • The HDF spec is too complex to write a simple read program... • ...without re-creating much of the API What to do?
HDF Mapping Files Concept: create text-based "maps” (cf. Mealy!!) of the HDF-4 file layouts while we still have a viable HDF-4 API (i.e., now) • XML • Stored separately from, but close to the data files • Includes • internal metadata • variable info • chunk-level info • byte offsets and length • linked blocks • compression information Task funded by ESDIS project • The HDF Group, NSIDC and GES DISC In XML -> ASCII format!
Map sample (extract) <hdf4:SDS objName="TotalCounts_A" objPath="/ascending/Data Fields" objID="xid-DFTAG_NDG-5"> <hdf4:Attribute name="_FillValue" ntDesc="16-bit signed integer"> 0 0 </hdf4:Attribute> <hdf4:Datatype dtypeClass="INT" dtypeSize="2" byteOrder="BE" /> <hdf4:Dataspace ndims="2"> 180 360 </hdf4:Dataspace> <hdf4:Datablock nblocks="1"> <hdf4:Block offset="27266625" nbytes="20582" compression="coder_type=DEFLATE" /> </hdf4:Datablock> </hdf4:SDS>
Their plans were…. (alas no funds) Status • Map creation utility (part of HDF) • Prototype read programs • C • Perl • Paper in TGRS special issue (a Journal) • Inventory of HDF-4 data products within EOSDIS (NASA data system) Future Steps • Revise XML schema • Revise map utility and add to HDF baseline • Implement map creation and storage operationally • e.g., add to ECS or S4PA metadata files (conventions)
But to really preserve? • CONTEXT!
NASA/ MODIS Contextual Info 7th Joint ESDSWG meeting, October 22, Philadelphia, PA Data Lifecycle Workshop sponsored by the Technology Infusion Working Group
Instrument/sensor characteristics Presented by R. Duerr at the Summer Institute on Data Curation, June 2-5, 2008 Graduate School of Library and Information Science, University of Illinois at Urbana-Champaign 46 7th Joint ESDSWG meeting, October 22, Philadelphia, PA Data Lifecycle Workshop sponsored by the Technology Infusion Working Group
Processing Algorithms & Scientific Basis Presented by R. Duerr at the Summer Institute on Data Curation, June 2-5, 2008 Graduate School of Library and Information Science, University of Illinois at Urbana-Champaign 47 7th Joint ESDSWG meeting, October 22, Philadelphia, PA Data Lifecycle Workshop sponsored by the Technology Infusion Working Group
Ancillary Data Presented by R. Duerr at the Summer Institute on Data Curation, June 2-5, 2008 Graduate School of Library and Information Science, University of Illinois at Urbana-Champaign 48 7th Joint ESDSWG meeting, October 22, Philadelphia, PA Data Lifecycle Workshop sponsored by the Technology Infusion Working Group
Processing History including Source Code Presented by R. Duerr at the Summer Institute on Data Curation, June 2-5, 2008 Graduate School of Library and Information Science, University of Illinois at Urbana-Champaign 49 7th Joint ESDSWG meeting, October 22, Philadelphia, PA Data Lifecycle Workshop sponsored by the Technology Infusion Working Group
Quality Assessment Information Presented by R. Duerr at the Summer Institute on Data Curation, June 2-5, 2008 Graduate School of Library and Information Science, University of Illinois at Urbana-Champaign 50 7th Joint ESDSWG meeting, October 22, Philadelphia, PA Data Lifecycle Workshop sponsored by the Technology Infusion Working Group