Transposable Elements
Transposable Elements. IS P-elements Human repetitive sequences. Prokaryotes. Insertion Sequences (IS elements) Composite transposons Tn3 elements. IS element. IS elements.

Transposable Elements
E N D
Presentation Transcript
Transposable Elements ISP-elements Human repetitive sequences
Prokaryotes • Insertion Sequences (IS elements) • Composite transposons • Tn3 elements
IS elements • IS elements are relatively small transposable elements that range in size from 760 to less than 2,500 base pairs (bp). • can insert at many different sites in bacterial and viral chromosomes and plasmids, and they contain genes whose products are involved in promoting and regulating transposition. • One of the genes is a transposase that functions in excision of the element from a chromosome, plasmid.
IS elements • IS elements typically generate unstable mutants that revert to wild-type at a detectable frequency. For that reason, IS elements originally were called "mutable" genes.
IS elements • All IS elements contain inverted terminal repeats that range in size (length) from 9 to 40 base pairs. • At the site of integration there invariably is a target site duplication of from 2‑13 base pairs.
Composite Transposons • Composite transposons (denoted by symbol Tn): • Tn elements stem from two IS elements that insert near one other. The regions (sequences) between the two elements can be "mobilized" by the joint action of the two IS elements. This is of significance in that many Tn elements possess genes that confer resistance to antibiotics between the two IS elements. • Tn transposition is regulated by a "repressor" that appears to exist to keep the elements somewhat quiescent.
Tn3 • Tn3 elements are simply large transposable elements that are not generated by flanking IS elements (as in Tn elements).
Eukaryotes • DNA transposable elements • RNA transposable elements
Conservative Transposition http://nitro.biosci.arizona.edu/courses/EEB600A-2003/lectures/lecture26/lecture26.html
AC/DS elements in maize • AC is a full-length autonomous copy • DS is a truncated copy of AC that is non-autonomous, requiring AC in order to transpose
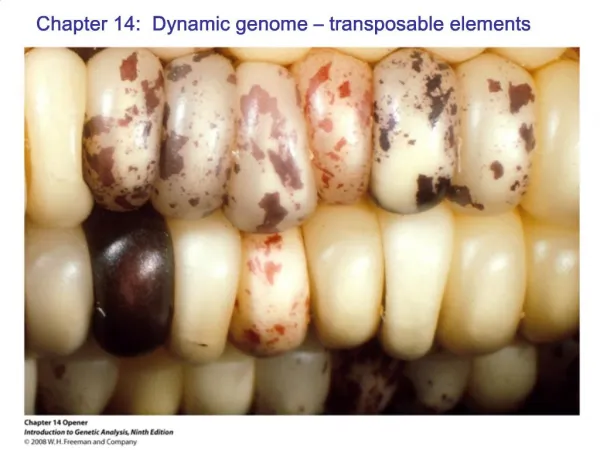
Transposable Elements (Transposons) • DNA elements capable of moving ("transposing") about the genome • Discovered by Barbara McClintock, largely from cytogenetic studies in maize, but since found in most organisms • She was studying "variegation" or sectoring in leaves and seeds • She liked to call them "controlling elements“ because they affected gene expression in myriad ways
Mutant Kernel Phenotypes • Pigmentation mutants • affect anthocyanin pathway • elements jump in/out of transcription factor genes (C or R) • sectoring phenotype - somatic mutations • whole kernel effected - germ line mutation
Some maize phenotypes caused by transposable elements excising in somatic tissues. Start with mutant kernels defective in anthocyanin synthesis and the element excises during development.
Somatic Excision of Ds from C Wild type Mutant Sectoring
Molecular Analysis of Transposons • Transposons isolated by first cloning a gene that they invaded. A number have been cloned this way, via "Transposon trapping“. • Some common molecular features: • Exist as multiple copies in the genome • Insertion site of element does not have extensive homology to the transposon • Termini are an inverted repeat • Encode “transposases” that promote movement • A short, direct repeat of genomic DNA often flanks the transposon : “Footprint”
Ac and Ds • Ds is derived from Ac by internal deletions • Ds is not autonomous, requires Ac to move • Element termini are an imperfect IR • Ac encodes a protein that promotes movement - Transposase • Transposase excises element at IR, and also cuts the target
Structure of Ac and Ds deletion derivatives Ds is not autonomous, requires Ac to move! Fig. 23.11
How duplications in the target site probably occur. Duplication remains when element excises, thus the Footprint.
Hybrid Dysgenesis • P-elements are transposable elements that carry genes for transposase activity that cause the elements to move, and repressor activity that prevents expression of transposase. • In a cross between a P-element-carrying female and a laboratory male [left], repressors in the maternally - derived cytoplasm repress expression of the maternally - inherited P elements. The resulting offspring show the wild-type phenotype.
Examples: P-elements in Drosophila • P elements were discovered when it was found that certain strains of Drosophila exhibited an assortment of aberrant phenotypes, including elevated mutation (and reversion), chromosome breakage, and sterility = hybrid dysgenesis • normally (within populations) the P elements are quiescent and do not “jump.” When “hybrids” were made between individuals from different geographic populations, the elements “moved” and promoted the dysgenic phenotypes.
P-elements • P elements vary in size (the largest are nearly 3,000 base pairs in length). Complete (intact) P elements possess a gene for a transposase. The number of P elements per individual varies from a few to up to 50.
Experimental uses of P-elements • Transposon tagging, where genes mutated by P element insertion can be isolated and "discovered" by using the P element sequence as a "tag”; and • Transformation vectoring, where genes or sequences of interest are "vectored" into a chromosomal location by putting the gene/sequence of interest into an incomplete P element (no transposase) and carrying out a mixed infection (transformation or electroporation) with a complete P element.
Mariner elements • Mariner elements appear to be a fairly widespread transposon of roughly 1,200 base pairs.
Retrotransposones • Retrovirus-like elements • Retroposons
LTR=long terminal repeat • Flanks three genes; a complete retrovirus has three genes • gag = structural gene for capsid • Pol = reverse transcriptase plus other stuff. • env = envelope gene for the virus
Retrovirus like elements • The basic structure of retrovirus-like elements is a central coding region of two genes flanked by long terminal repeats [LTRs] that are oriented in the same direction and bounded by short inverted repeats. • The two genes are homologous to two genes in retroviruses and encode a structural protein of the virus capsule and a reverse transcriptase/integrase enzyme.
Retrovirus like elements • Active retroviruses carry a third gene that codes for a protein of the virus envelope. Active retroviruses are capable of exiting cells and infecting other cells. • Transposition involves transcription (RNA synthesis) of the DNA sequence integrated in the chromosome, reverse transcription of the RNA, synthesis of a double-stranded DNA from the RNA, and insertion into a new chromosomal location.
Retroposons • These are elements that move through an RNA intermediary but do not possess direct or inverted repeats at their termini (LTR) nor the env gene. • They possess instead a string of A=T base pairs at one end (of the DNA), and presumably represent a copy from reverse transcription of the poly-A tail of the mature RNA transcript. • Some LINE sequences in mammals are retroposons, and the LlNE‑1 retroposon is the only transposable element thus far documented in humans. • Drosophila telomere sequences
Non-LTR transposition The LINE is transcribed into mRNA (red). A part of this mRNA is translated into proteins involved in the integration complex, which binds to the 3' end of the mRNA transcript. The target site (blue) is cleaved followed by reverse transcription, with the 3' end of the target site as the primer. Newly synthesized cDNA is shown in pale green. Ligation of the cDNA occurs at the 5' end, and the second strand is synthesized using the first cDNA strand as template and the host DNA polymerase
Long term fate of non-LTR • In an evolving genome, non-LTR elements are thought to proliferate by amplification of an extremely small number of "master" genes. • These genes usually give rise to inactive copies (truncated at the 5' end) that are incapable of further transposition within the genome. • The defective copies arise because of their mode of transposition through reverse transcription (see the figure), which in most cases stops replication before the 5' end is reached. • These truncated elements, called DOA ("dead on arrival"), can be used as surrogates for pseudogenes in species such as Drosophila that have few bona fide pseudogenes
LINEs • In humans, are about 6 kb long, harbour an internal polymerase II promoter and encode two open reading frames (ORFs). • Upon translation, a LINE RNA assembles with its own encoded proteins and moves to the nucleus, where an endonuclease activity makes a single-stranded nick and the reverse transcriptase uses the nicked DNA to prime reverse transcription from the 3' end of the LINE RNA. • Reverse transcription frequently fails to proceed to the 5' end, resulting in many truncated, nonfunctional insertions.
LINEs • Most LINE-derived repeats are short, with an average size of 900 bp for all LINE1 copies, and a median size of 1,070 bp for copies of the currently active LINE1 element (L1Hs). • The LINE machinery is believed to be responsible for most reverse transcription in the genome, including the retrotransposition of the non-autonomous SINEs and the creation of processed pseudogenes • Three distantly related LINE families are found in the human genome: LINE1, LINE2 and LINE3. • Only LINE1 is still active.
SINEs • SINEs are freeloaders on the backs of LINE elements. • short (about 100-400 bp), harbour an internal polymerase III promoter and encode no proteins. • non-autonomous transposons use the LINE machinery for transposition.
SINEs • most SINEs 'live' by sharing the 3' end with a resident LINE element. • Most promoter regions of known SINEs are derived from tRNA sequences
Alu • A single monophyletic family of SINEs (ALU) derived from the signal recognition particle component 7SL • This family is the only active SINE in the human genome • The human genome contains three distinct monophyletic families of SINEs: the active Alu, and the inactive MIR and Ther2/MIR3.
Direct Repeats • duplication of a short sequence at the target site. This generates short direct repeats flanking the newly inserted element. This results for a staggered cut being made in the DNA strands at the site of insertion
LTR retroposons • Bascially, these are retroviruses without the env protein. • Current thinking is that retroviruses evolved from retroposons. • They have the LTR and (usually) gag genes. LTR retroposons are often simple called retrotransposons.