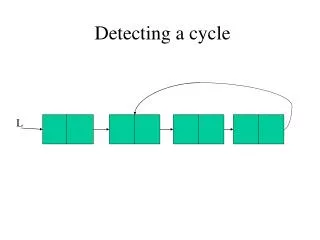
A Method for Detecting Pleiotropy
This method tests for independent genetic effects on two traits, estimating the parameter of pleiotropy and comparing null and alternative hypotheses using MANOVA, FCP, RCM, and SUM tests.


A Method for Detecting Pleiotropy
E N D
Presentation Transcript
A Method for Detecting Pleiotropy Ingrid Borecki, Qunyuan Zhang, Michael Province Division of Statistical Genomics Washington University School of Medicine
Biological question: Does a genetic variant have independent effects on both of two traits? Pleiotropy Statistical question: Can the correlation or a portion of the correlation between two traits be explained by a genetic variant?
Compound null: no pleiotropy Alternative: pleiotropy Hypotheses & Models Y1 Y2 X Y1 Y2 Y1 Y2 Y1 Y2 X X X
Statistical Parameter (δ) of Pleiotropy& Hypotheses to Be Tested Compound null: no pleiotropy Alternative: pleiotropy
Estimating δ Two traits are simultaneously fit into a mixed model T is the trait indicating variable; R is block diagonal covariance matrix (after re-ordering by individuals), with blocks corresponding to the individuals and each block having the compound-symmetry structure When excluding X from the model When including X in the model
Q-Q Plot under the null Testing δ Pleiotropy Estimation Test (PET) Estimated by bootstrapre-sampling 100 times with replacement -LOG10(P)
=Residual of Y1 adjusted by Y2 =Residual of Y2 adjusted by Y1 • MANOVA (Wilks' test, wrong null) • FCP: Fisher’s combined p-value test (meta-analysis ignoring correlations, wrong null) • RCM: Reverse compound model (two tests) • SUM: Simple univariate model (two tests) Other Methods for Comparison Testing if β1≠0 and β2≠0
Power Comparison PET FCP MANOVA RCM SUM
Power Comparison PET FCP MANOVA RCM SUM
Correlation (WC, HOMA)= 0.542 Application Correlation (TG, CAC)= 0.089
The PET Method • Tests proper compound null for pleiotropy; • Gives estimation of covariance due to pleiotropy; • Has greater power other alternatives; • Under mixed model framework, can easily be expanded to other data (covariates, family data etc.) ; • Practical to GWAS data (with 300 blades, R version takes less than 1 day for the analysis of 2M SNPs and ~3000 subjects) ; • Must be fit to primary phenotype and (typed or imputed) genotype data. Conclusions
Acknowledgement Ling-Yun Chang (programming & testing) Mary Feitosa (GWAS data and application)