Comparative Analysis of ISE2 and Ski2P RNA Helicase Motifs in Diverse Organisms
This study presents a comparative analysis of the consensus DEVH RNA helicase motifs in ISE2 and Ski2P from yeast. The findings include a phylogenetic tree illustrating the homologs of ISE2 across various organisms, created using the neighbour-joining method with the BLOSUM 62 matrix. The analysis highlights the evolutionary distances between different ISE2 homologs, emphasizing their significance in molecular biology. This research is adapted from the work by Kyle Tanner and Patrick Linder published in Molecular Cell.

Comparative Analysis of ISE2 and Ski2P RNA Helicase Motifs in Diverse Organisms
E N D
Presentation Transcript
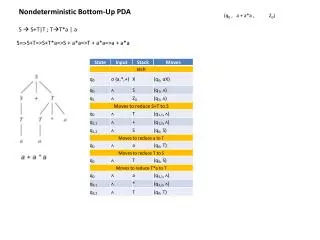
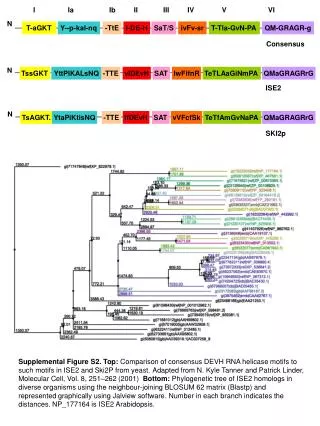
I Ia Ib II III IV V VI N T-aGKT Y--p-kal-nq -TtE I-DE-H SaT/S ivFv-sr T-Tla-GvN-PA QM-GRAGR-g Consensus N TssGKT YttPlKALsNQ -TTE vlDEvH SAT IwFifnR TeTLAaGiNmPA QMaGRAGRrG ISE2 N TsAGKT. YtaPiKtisNQ -TTE IfDEvH SAT vVFcfSk TeTfAmGvNaPA QMaGRAGRrG SKI2p Supplemental Figure S2. Top: Comparison of consensus DEVH RNA helicase motifs to such motifs in ISE2 and Ski2P from yeast.Adapted from N. Kyle Tanner and Patrick Linder, Molecular Cell, Vol. 8, 251–262 (2001) Bottom: Phylogenetic tree of ISE2 homologs in diverse organisms using the neighbour-joining BLOSUM 62 matrix (Blastp) and represented graphically using Jalview software. Number in each branch indicates the distances. NP_177164 is ISE2 Arabidopsis.






![[ t-t-t ]](https://cdn1.slideserve.com/3099327/slide1-dt.jpg)