Understanding DNA Replication: A Semi-conservative Process Explained
DNA replication is a crucial biological process where the DNA molecule is duplicated, ensuring genetic continuity. This semi-conservative process involves several steps and proteins at origins of replication, forming replication bubbles and forks. Key enzymes like helicase unwound the DNA, while DNA polymerases synthesize new strands from RNA primers. The leading and lagging strands highlight the antiparallel nature of DNA, with the latter synthesized in fragments. Understanding these mechanisms provides insights into how your body maintains its genetic information day by day.

Understanding DNA Replication: A Semi-conservative Process Explained
E N D
Presentation Transcript
Review • Why is the process of DNA called a semiconservative process? • Let’s review DNA structure: • List all characteristics you can recall about DNA molecule
A closer look • Today we will learn about some incredible stuff happening in your body right now! (p.313)
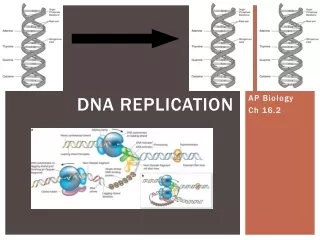
Getting Started • DNA replication begins @ particular sites called origins of replication • Proteins attach and begin copying DNA strand • Forms replication bubble could be multiple in single DNA helix = speeds up replication process • At each end of bubble = replication forks: the y=shaped region where parts of parental DNA being unwound DRAW pic. on pg. 313
Proteins involved in unwinding • Helicase: untwists/unzips DNA at forks =opens up template strands • Single –strand binding proteins: bind to unwound parental strands, keeping them from re-pairing • Topoisomerase: helps relieve strain on unwound DNA while unwinding is occurring so DNA does not break permanently
Replication Initiation • Enzymes that add nucleotides need help to start… can’t initiate process on their own • initial nucleotide chain is added = a short RNA chain called a primer created by enzyme called primase -usually 5-10 nucleotides long *new DNA strand will start from 3’ end of primer
Adding Bases • Main protein involved = DNA polymerase Specifically DNA polymerase I (DNA pol I) and DNA polymerase III (DNA pol III) • After primer added, polymerase will begin adding nucleoside triphosphates (dATP) to the strand • Similar to ATP except sugar: ATP = ribose, dATP = deoxyribose When added to template, 2 phosphates break off (b/c unstable) = exergonic reaction that drives polymerization
Antiparallel Elongation • DNA strands/replication has directionality (like 1 way street) compliment strands are antiparallel How does the antiparallel arrangement of the double helix affect replication? (HINT: think back to the rule of the primer)
Answer: • DNA pol can only add bases to 3’ end of primer .. So a new DNA strand can ONLY elongate from 5’-3’ direction Let’s take a closer look at replication forks of the bubble to see how this works!
Antiparallel Elongation • 1st strand, DNA pol III can synthesize complimentary strand continuously as DNA unwinds from 5’ – 3’ direction = LEADING STRAND • Only 1 primer needed • 2nd strand, DNA pol III must work in opposite direction (away from replication fork) to continue replication in 5’ – 3’ direction = LAGGING STRAND Synthesized discontinuously in segments called Okazaki fragments
What do we do with the primer? • When segment replication is complete, primer must be removed • DNA pol I comes in and removes RNA primer and fills hole with DNA nucleotides • An enzyme called DNA ligase comes in and patches two segments of lagging strand together Let’s see it in action!
Review: • For the following enzymes/proteins, list their roles in DNA replication: • DNA polymerase III • DNA helicase • DNA ligase • Primase • Single-strand binding proteins • Topoisomerase • DNA ligase