Extracting and Exploiting Structural Patterns in Proteins, especially Relating to Function
Extracting and Exploiting Structural Patterns in Proteins, especially Relating to Function. Janet Thornton James Watson, Roman Laskowski - EBI Adel Golovin, Kim Henrick - EBI MSD David Leader, James Milner-White – Glasgow Andrzej Joachimiak, Aled Edwards – MCSG

Extracting and Exploiting Structural Patterns in Proteins, especially Relating to Function
E N D
Presentation Transcript
Extracting and Exploiting Structural Patterns in Proteins, especially Relating to Function Janet Thornton James Watson, Roman Laskowski - EBI Adel Golovin, Kim Henrick - EBI MSD David Leader, James Milner-White – Glasgow Andrzej Joachimiak, Aled Edwards – MCSG (Mid-West Centre for Structural Genomics)
Outline • Structural Motifs • PDBsum • MSDmotif • Functional Motifs • Catalytic Site Atlas • DNA Binding Motifs • Automated templates • Reverse Templates • From Structure to Function? - ProFunc
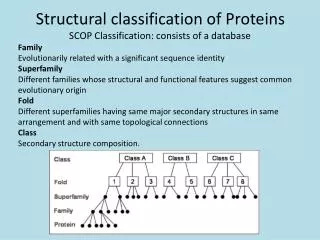
Structural Motifs Structural motifs are commonly occurring small sections of proteins – that are distinguished by: Sequence – Gly-X-Gly Conformation – , angles Secondary structure - helix, bab unit Function – catalytic triad, calcium binding site
Examples of Structural Motifs AlphaBeta Motif Beta Turn Schellmann Loop Beta Bulge (classic) Nest Beta Bulge Loop
Structural Motifs They may be continuous along the chain (e.g. GXG) or discontinuous (e.g. catalytic triad) Historically motifs were identified and analysed in an effort to understand the relationship between protein sequence and structure, to improve prediction methods. They are also used to assign function (Prosite). Many motifs can now be recognised automatically from coordinates, using programmes such as DSSP and Promotif PDB files can be annotated with these structural motifs e.g. in PDBsum
http://www.ebi.ac.uk/thornton-srv/databases/pdbsum/ Roman Laskowski
MSD motifhttp://www.ebi.ac.uk/msd-srv/msdmotif Adel Golovin Currently alpha test Full Release probably ~Oct 2005 PDB: 1gci
MSD motif Small 3D motifs from J.Milner-White search/view Secondary structure patterns (HTH) search/view ,, based search/view Ligands and their environment search/view Catalytic sites search/view Blast sequence search/view Prosite compliant patterns search/view 3D multiple alignment
Small motifs Alpha-Beta Motif Nest ST staple 11 motifs in total (Prof James Milner-White) http://doolittle.ibls.gla.ac.uk:9006/david/ProteinMotifDB.html
Motifs In MSDmotif (1) AlphaBeta Motif Beta Turn Schellmann Loop Beta Bulge (classic) Nest Beta Bulge Loop
Motifs In MSDmotif (2) Asx Motif ST Motif Asx Turn ST Turn ST Staple
Statistics provided by MSDmotifSTmotif a) b) c) • Amino acid occurrence at each position • Correlation between side chain charge and residue position • Motif parameter variation
Small motifs – 3D alignmentfrom different families ST-staple
Strand – turn – Strand 2-3 residues gap Glycosylation pattern N{P}[ST]{P} Secondary structure patterns Where N binds sugar: Man or Nag
,, search PDB:1gci Ideal for short loops search
Example of a search using MSDmotif PDB:1gci Subtilases family PDB:1f5p Globins family Phi/Psi Search using MSDmotif + Other Subtilases Calcium binding site
Sequence search ZN binding pattern: CXXCXXXFXXXXXLXXHXXXH
MSD motif • Available in alpha version • http://www.ebi.ac.uk/msd-srv/msdmotif • Will be published later this year • Incremental weekly update • 20 G disk space on Oracle DB, linear dependency ~ 0.8 M per PDB • Web application server with J2EE servlet engine • NCBI Blast Adel Golovin Kim Henrick
Outline • Structural Motifs • PDBsum • MSDmotif • Functional Motifs • Catalytic Site Atlas • DNA Binding Motifs • Automated templates • Reverse Templates • From Structure to Function? - ProFunc
Catalytic Site Atlas • Taken from primary literature: • -lactamase Class A • EC: 3.5.2.6 • PDB: 1btl • Reaction: -lactam + H2O -amino acid • Active site residues: S70, K73, S130, E166 • Plausible mechanism:
The Catalytic Site Atlas: a resource of catalytic sites and residues identified in enzymes using structural data. Craig T. Porter, Gail J. Bartlett, and Janet M. Thornton Nucl. Acids. Res. 2004 32: D129-D133. http://www.ebi.ac.uk/thornton-srv/databases/CSA
Annotates catalytic residues in the PDB • Based on a dataset of 514 enzyme families • Representative catalytic site for each family • Homologues assigned by Psi-BLAST • Limited substitution allowed. • Homologues updated monthly. • Literature references • Data also available via MSDsite • http://www.ebi.ac.uk/thornton-srv/databases/CSA • http://www.ebi.ac.uk/msd-srv/msdsite
3-D templates • Use 3D templates to describe the active site of the enzyme • analogous to 1-D sequence motifs such as PROSITE, butin 3-D • Sequence position independent • Captures essence of functional site in protein
Aspartic Proteinase - Active Site residues - [DTG]x2 Eukaryotic & Fungal Aspartic Proteinases: all-atom DTG-DTG Template
Aspartic Proteases: Active Site Template Asp CO2 Gly C A template of 8 atoms is sufficient to identify all Aspartic Proteinases Asp O Gly C Thr/Ser O Thr O
Aspartic Protease Template Search against all PDB green= true red=false
TEmplate Search and Superposition TESS Wallace et al., 1997 • defines a functional site as a sequence-independent set of atoms in 3-D space • search a new structure for a functional site • search a database of structures for similar clusters e.g. serine proteinase, catalytic triad
Serine Proteinase templates • A trypsin-based template of 7 atoms was able to identify almost all serine proteinases in PDB- including subtilisin • It also identified active sites of several other functionally distinct enzyme families - serine carboxypeptidase, acetylcholine esterase; lipase; dehalogenase • The catalytic triad has evolved independently many times
Active site convergence Trypsin Subtilisin
Trypsin Subtilisin Alpha/beta hydrolase Brain platelet activating factor acetylhydrolase Clp protease CheB methylesterase
(~600 Metal binding site templates) (189 enzyme active site templates) 3D Templates to Characterise Functional Sites Template searches
GARTfase Cholesterol oxidase IIAglc histidine kinase Database of enzyme active site templates 189 templates … Carbamoylsarcosine amidohhydrase Ser-His-Asp catalytic triad Dihydrofolate reductase
DNA Protein +
DNA-binding Motifs • Helix-Turn-Helix (HTH) • Standard HTH • Winged helix • Beta Sheet • Zinc-finger
Prediction of DNA Binding Function using Structural Motifs • Predicting function from structure • Structural motifs • Helix-Turn-Helix (HTH) • Bind in major groove • Carboxyl terminal helix - DNA recognition • 1/3 DNA-binding protein families (16/54) • Brennan and Mathews 1989: Brennan, 1991
HTH Motif Proteins Catabolic activator protein (1ber) Lambda repressor/operator complex (1lmb)
HTH Motif Templates 3D template library (E.g. 1berA16-36)
Predicting DNA binding function • Scanning template library against 3D structures • One templateT(length n) scanned against proteinP of length m, RMSD calculated optimal superposition at each m-n+1 possible positions in P • Calculate lowest RMSD for optimal superposition
RMSD Distributions with HTH templates 1.2Å RMSD 831/23,506 = 3.5% false positives 2/142 = 1.4% false negatives
HTH Motif Extended Templates • Extend templates by adding +2 residues to start and end • 1berA16-36 • 1berA14-38
RMSD Distributions with extended HTH templates 1.2Å 110/23,506 = 0.5% false positives 2/144 = 1.4% false negatives