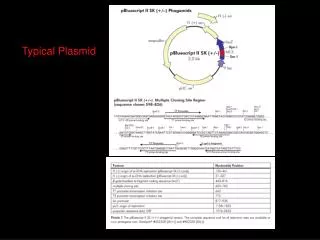
Typical Plasmid
Typical Plasmid. Blue/White Selection. Alpha complementation Trick. omega. alpha. Blue/White Selection. Standard 4 exon gene. -Where is the polyA tail in the gene above? -Draw a 4 exon gene where exon 4 is all 3’UTR. Chapter 11 Transcription and RNA Processing. The Central Dogma.

Typical Plasmid
E N D
Presentation Transcript
Alpha complementation Trick omega alpha
Standard 4 exon gene -Where is the polyA tail in the gene above? -Draw a 4 exon gene where exon 4 is all 3’UTR
Transcription and Translation in Prokaryotes • The primary transcript is equivalent to the mRNA molecule. • The mRNA codons on the mRNA are translated into an amino acid sequence by the ribosomes.
Transcription and Translation in Eukaryotes • The primary transcript (pre-mRNA) is a precursor to the mRNA. • The pre-mRNA is modified at both ends, and introns are removed to produce the mRNA. • After processing, the mRNA is exported to the cytoplasm for translation by ribosomes.
Types of RNA Molecules • Messenger RNAs (mRNAs)—intermediates that carry genetic information from DNA to the ribosomes. • Transfer RNAs (tRNAs)—adaptors between amino acids and the codons in mRNA. • Ribosomal RNAs (rRNAs)—structural and catalytic components of ribosomes. • Small nuclear RNAs (snRNAs)—structural components of spliceosomes. • Micro RNAs (miRNAs)—short single-stranded RNAs that block expression of complementary mRNAs.
RNA Synthesis And Transport in Eukaryotes • Method: Pulse-Chase Labeling • At first, labeled RNA is exclusively in the nucleus. • Later, the labeled RNA is found in the cytoplasm.
General Features of RNA Synthesis • Similar to DNA Synthesis except • The precursors are ribonucleoside triphosphates ATP, GTP, CTP, UTP. • Only one strand of DNA is used as a template. • RNA chains can be initiated de novo (no primer required). • The RNA molecule will be complementary to the DNA template (antisense) strand and identical (except that uridine replaces thymidine) to the DNA nontemplate (sense) strand. • RNA synthesis is catalyzed by RNA polymerases and proceeds in the 5’3’ direction.
Prokaryotes-transcription and translation occur at the same time Transcription—the first step in gene expression—transfers the genetic information stored in DNA—genes—into messenger RNA molecules that carry the information to the ribosomes—the sites of protein synthesis—in the cytoplasm.
E. Coli RNA Polymerase • Tetrameric core: 2’ • Holoenzyme: 2’ • Functions of the subunits: • : assembly of the tetrameric core • : ribonucleoside triphosphate binding site • ’: DNA template binding region • : initiation of transcription
Initiation of RNA Chains • Binding of RNA polymerase holoenzyme to a promoter region in DNA • Localized unwinding of the two strands of DNA by RNA polymerase to provide a single-stranded template • Formation of phosphodiester bonds between the first few ribonucleotides in the nascent RNA chain
Numbering of a Transcription Unit • The transcript initiation site is +1. • Bases preceding the initiation site are given minus (–) prefixes and are referred to as upstream sequences. • Bases following the initiation site are given plus (+) prefixes and are referred to as downstream sequences.
A Typical E. coli Promoter • Consensus sequences: -10 sequence and -35 sequence • Recognition sequence: -35 sequence
Termination Signals in E. coli • Rho-dependent terminators—require a protein factor () • Rho-independent terminators—do not require
Transcription and RNA Processing in Eukaryotes Three different enzymes catalyze transcription in eukaryotes, and the resulting RNA transcripts undergo three important modifications, including the excision of noncoding sequences called introns. The nucleotide sequenced of some RNA transcripts are modified posttranscriptionally by RNA editing.
Modifications to Eukaryotic pre-mRNAs • A 7-Methyl guanosine cap is added to the 5’ end of the primary transcript by a 5’-5’ phosphate linkage. • A poly(A) tail (a 20-200 nucleotide polyadenosine tract) is added to the 3’ end of the transcript. The 3’ end is generated by cleavage rather than by termination. • When present, intron sequences are spliced out of the transcript.
Eukaryotes Have Three RNA Polymerases Pol II is the only Polymerase that is routinely studied. Pol I and Pol III are very complicated.
The 3’ Poly(A) Tail AATAAA
Interrupted Genes in Eukaryotes: Exons and Introns Most eukaryotic genes contain noncoding sequences called introns that interrupt the coding sequences, or exons. The introns are excised from the RNA transcripts prior to their transport to the cytoplasm.
Removal of Intron Sequences by RNA Splicing The noncoding introns are excised from gene transcripts by several different mechanisms.
Splicing • Removal of introns must be very precise. • Conserved sequences for removal of the introns of nuclear mRNA genes are minimal. • Dinucleotide sequences at the 5’ and 3’ ends of introns. • An A residue about 30 nucleotides upstream from the 3’ splice site is needed for lariat formation.
Types of Intron Excision • The introns of tRNA precursors are excised by precise endonucleolytic cleavage and ligation reactions catalyzed by special splicing endonuclease and ligase activities. • The introns of nuclear pre-mRNA (hnRNA) transcripts are spliced out in two-step reactions carried out by spliceosomes.
The Spliceosome • Five snRNAs: U1, U2, U4, U5, and U6 • Some snRNAs associate with proteins to form snRNAs (small nuclear ribonucleoproteins)