Domains
Domains. typical size ~100-200 amino acids mean=160 residues balance surface area to volume (hydrophobics in core) modularity, though insertions are possible a, b, a+b, a/b ( “wound” bab, parallel strands ) classic folds: globins, immunoglobulins, TIM-barrels, NBDs

Domains
E N D
Presentation Transcript
Domains • typical size ~100-200 amino acids • mean=160 residues • balance surface area to volume (hydrophobics in core) • modularity, though insertions are possible • a, b, a+b, a/b (“wound” bab, parallel strands) • classic folds: globins, immunoglobulins, TIM-barrels, NBDs • beta-sandwiches/clamshells, helix-bundles • helical coiled-coils (collagen, lambda repressor) • beta-barrels, beta-propellers
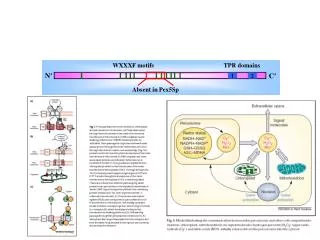
active site in mouth of beta-barrel • domain insertions • (the strange case of malate synthase...) C-terminal cap see 4 domains on CATH page: http://www.cathdb.info/chain/1n8wA
How small can a protein be and still have structure? • no hydrophobic core • glucagon (30 res, a-helix); dis-ordered in solution • unraveling, conformational sampling • NMR studies of peptides? • 10-aa SCF recognition peptide • disorder of p53 fragment in soln by NMR • on the contrary, 17-residue fragment from N-terminal domain of ubiquitin folds into beta-hairpin on its own • Zarella et al, Protein Science, 1999
Structure Superposition Algorithms • least-squares • Aij =∑PkiQkj– product over 2 sets of coords, P and Q • R = (AtA)1 / 2A−1 – rotation that minimizes RMSD • assumes translated to centers-of-mass • Kabsch rotation algorithm (1976, Acta) – (equiv. to SVD) • MacKay (1984, Acta) – quaternions (solve linear system) • SSAP (Orengo and Taylor) • dynamic programming to minimize inter-molecular distance vectors between Cb atoms • pairs must be known a priori let mij be eigenvalues and aij be eigenvectors of RTR: Lij are Lagrange multipliers. determine by solving:
DALI (Holm and Sander) • aligns scalar distance plots • significance: z-scores: z=(s-m)/s>7.0 • compare to scores from random alignments • beware of effect of length of aligned/rejected; shorter->better score
VAST (Gibrat and Bryant) • aligns secondary structure elements • graph theory algotrithm – finds maximal clique in graph of consistent alignable pairs of vectors • LOCK (Singh and Brutlag) • hierarchical, distances + SS elements • rigid bodies can’t always be aligned well • CE (combinatorial extension; Shindalyov&Bourne) • identifies similar local fragments (3-5aa), extends them • more tolerant of flexible regions • SSM (Krissinel and Henrick) • subgraph isomorphism • must preserve topology?
Fold Families • clustering • PDBSelect and COG are based on homology only • FSSP - based on DALI score • SCOP – manually curated (by Alexy Muzrin) • CATH (Orengo and Thornton) • Pfam – based on HMMs (more details later)
(beware of large-family bias when averaging over protein database)
Fold Recognition • sequence alignment (homology) • position-dependent profiles from multiple alignment (Gribskov, McLachlan, Eisenberg, 1987), scores based on sum of Dayhoff similarity over observed residues at each pos. • 3D profiles • threading • HMMs
Convergence vs. Divergence Sander and Schneider (1991) Database of Homology-Derived Protein Structures and the Structural Meaning of Sequence Alignment. Chothia, C. (1993). One thousand families for the molecular biologist.
3D Profiles (Eisenberg et al.) • Given that you have a sequence threaded onto a known structure, how well does it fit the fold? • originally: residues scored by 18 environment classes (Bowie, Luthy, Eisenberg, 1991) • similarity of amino acids in model to structure (homology, position-dependent distribution) • tolerance of buried vs. surface exposure • suitability of residues in secondary structures • residue pair potentials (likelihood of contacts at 4-10A radius shells) (Wilmanns and Eisenberg, 1993)
18 environment classes = {E,P1,P2,B1,B2,B2}x{helix,sheet,coil}
Threading (for Fold Recognition) • find optimal mapping of residues in sequence to model • higher computational complexity that sequence alignment, or can also be done by dynamic programming? • Lathrop (Prot Eng, 1994; JMB, 1996) - showed that threading is NP-complete when non-local effects are taken into account (reduction to 3SAT) • fold evaluation: • 3D profiles • packing (steric conflicts, voids) • energy (molecular mechanics force field) • statistical (side-chain contacts, Sippl) • PHYRE (Sternberg) – 3D-PSSM search • THREADER (David Jones) – dynamic programming • RAPTOR (Jinbo Xu) – integer programming (constraints)
Pfam, Hidden Markov Models (HMMs) (Sonnhammer, Eddy, and Durbin, 1997) Viterbi algorithm (forward/backward) training: maximum likelihood, EM
HMM for 628 globins (lines indicate most frequently- used transitions)
1YBA – PDGH tetramer 1BEF -protease Linkers • definition: • do not pack against well-defined domains (lack contact; not necessarily exposed, though) • can’t count on sequence between known domains • flexible, lack regular secondary structure (not always coil; helical linkers exist) • rich in Pro, Ala, charged residues; lack of Gly • George and Heringa (2002) • Bae, Mallick, Elsik (2005) – HMM (accuracy ~ 67%) 4FAB - immunoglobulin
Tanaka, Yokayama, Kuroda (2006) – length dependence • significant frequency deviations were observed for glycine, proline, and aspartic acid in short linker and nonlinker loops, whereas deviations were observed for aspartic acid, proline, asparagine, and lysine in long linker and nonlinker loops. all fragments length <= 9 aa length > 9 aa • DomCut (Suyama & Ohara, 2003) • uses differences in amino acid composition between the intra- and interdomain regions to predict domain boundaries • Armadillo (Dumontier et al., 2005) • local smoothing of aa propensity index by FFT; calculates Z-score