Genomic Analysis of Smoking Behavior-Associated Genes in Nicotinic Acetylcholine Receptor Families
10 likes | 96 Vues
Explore SNPs in 3’UTR regions and their impact on miRNA binding, focusing on CHRNA and CHRNB families, CYP2A6, and DRD1. Investigate potential genetic modulators influencing smoking behavior.

Genomic Analysis of Smoking Behavior-Associated Genes in Nicotinic Acetylcholine Receptor Families
E N D
Presentation Transcript
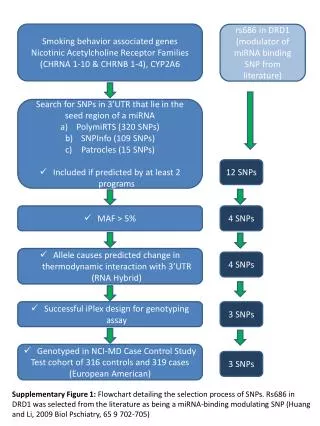
Smoking behavior associated genes Nicotinic Acetylcholine Receptor Families (CHRNA 1-10 & CHRNB 1-4), CYP2A6 rs686 in DRD1 (modulator of miRNA binding SNP from literature) • Search for SNPs in 3’UTR that lie in the seed region of a miRNA • PolymiRTS (320 SNPs) • SNPInfo (109 SNPs) • Patrocles (15 SNPs) • Included if predicted by at least 2 programs 12 SNPs • MAF > 5% 4 SNPs • Allele causes predicted change in thermodynamic interaction with 3’UTR (RNA Hybrid) 4 SNPs • Successful iPlex design for genotyping assay 3 SNPs • Genotyped in NCI-MD Case Control Study • Test cohort of 316 controls and 319 cases (European American) 3 SNPs Supplementary Figure 1: Flowchart detailing the selection process of SNPs. Rs686 in DRD1 was selected from the literature as being a miRNA-binding modulating SNP (Huang and Li, 2009 Biol Pschiatry, 65 9 702-705)