Nucleotide metabolism – Part 1 (purine biosynthesis)
Nucleotide metabolism – Part 1 (purine biosynthesis). By Henry Wormser, Ph.D Professor of Medicinal Chemistry. Biological significance of nucleotide metabolism. Nucleotides make up nucleic acids (DNA and RNA) Nucleotide triphosphates are the “energy carriers” in cells (primarily ATP)

Nucleotide metabolism – Part 1 (purine biosynthesis)
E N D
Presentation Transcript
Nucleotide metabolism – Part 1(purine biosynthesis) By Henry Wormser, Ph.D Professor of Medicinal Chemistry
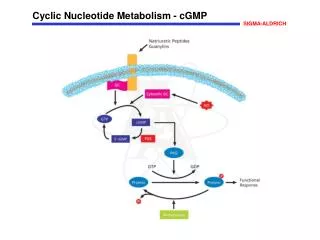
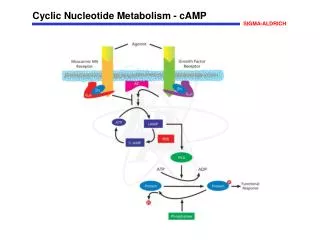
Biological significance of nucleotide metabolism • Nucleotides make up nucleic acids (DNA and RNA) • Nucleotide triphosphates are the “energy carriers” in cells (primarily ATP) • Many metabolic pathways are regulated by the level of the individual nucleotides • Example: cAMP regulation of glucose release • Adenine nucleotides are components of many of the coenzymes • Examples: NAD+, NADP+, FAD, FMN, coenzyme A
Dietary nucleotides • do not contribute energy as do carbs, proteins and fats • are not incorporated into RNA or DNA unless given I.V. • normally metabolized to individual components (bases, sugar and phosphate) • purines are converted to uric acid which is then excreted
Medical significance of nucleotide metabolism • Anticancer agents: • Rapidly dividing cells biosynthesize lots of purines and pyrimidines, but other cells reuse them. Cancer cells are rapidly dividing, so inhibitor of nucleotide metabolism kill them • Antiviral agents • Zidovudine (Retrovir) • Lamivudine (Epivir) • Valacyclovir (Valtrex)
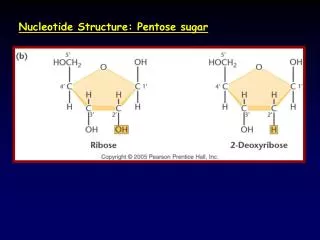
Structures of nucleotide buildingblocks and nucleotides guanine: comes from guano; thymine –thymus gland
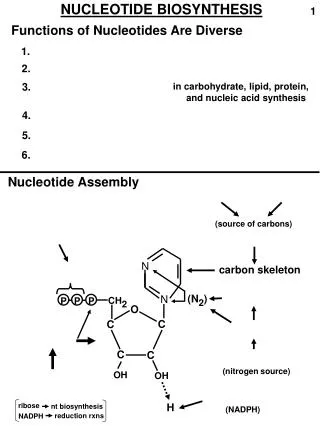
Synthesis of Inosine Monophosphate • Basic pathway for biosynthesis of purine ribonucleotides • Starts from ribose-5-phosphate which is derived from the pentose phosphate pathway • Requires 11 steps overall • occurs primarily in the liver
Steps 1 thru 3 • Step 1:Activation of ribose-5-phosphate • enzyme: ribose phosphate pyrophosphokinase • product: 5-phosphoribosyl-a-pyrophosphate (PRPP) • PRPP is also a precursor in the biosynthesis of pyrimidine nucleotides and the amino acids histidine and tryptophan
Steps 1 thru 3 • Step 2: acquisition of purine atom 9 • enzyme: amidophosphoribosyl transferase • displacement of pyrophosphate group by glutamine amide nitrogen (inversion of configuration – a to b • product: b-5-phosphoribosylamine Steps 1 and 2 are tightly regulated by feedback inhibition
Steps 1 thru 3 • Step 3: acquisition of purine atoms C4, C5, and N7 • enzyme: glycinamide synthetase • b-phosphoribosylamine reacts with ATP and glycine • product: glycinamide ribotide (GAR)
Steps 4 thru 6 • Step 4: acquisition of purine atom C8 • formylation of free a-amino group of GAR • enzyme: GAR transformylase • co-factor of enzyme is N10-formyl THF • Step 5: acquisition of purine atom N3 • The amide amino group of a second glutamine is transferred to form formylglycinamidine ribotide (FGAM) • Step 6: closing of the imidazole ring or formation of 5-aminoimidazole ribotide
Step 7 • Step 7: acquisition of C6 • C6 is introduced as HCO3- • enzyme: AIR carboxylase (aminoimidazole ribotide carboxylase) • product: CAIR (carboxyaminoimidazole ribotide) • enzyme composed of 2 proteins: PurE and PurK (synergistic proteins)
Steps 8 thru 11 • Step 8: acquisition of N1 • N1 is acquired from aspartate in an amide condensation reaction • enzyme: SAICAR synthetase • product: 5-aminoimidazole-4-(N-succinylocarboxamide)ribotide (SAICAR) • reaction is driven by hydrolysis of ATP
Steps 8 thru 11 • Step 9: elimination of fumarate • Enzyme: adenylosuccinate lyase • Product: 5-aminoimidazole-4-carboxamide ribotide (AICAR) • Step 10: acquisition of C2 • Another formylation reaction catalyzed by AICAR transformylase • Product: 5-formaminoimidazole-4-carboxamide ribotide (FAICAR)
Step 11 • cyclization or ring closure to form IMP • water is eliminated • in contrast to step 6 (closure of the imidazole ring), this reaction does not require ATP hydrolysis • once formed, IMP is rapidly converted to AMP and GMP (it does not accumulate in cells
Synthesis of adenine and guanine nucleotides
Purine nucleoside diphosphates and triphosphates: - to be incorporated into DNA and RNA, nucleoside monophosphates (NMP’s) must be converted into nucleoside triphosphates (NTP’s) - nucleoside monophosphate kinases (adenylate & guanylate kinases) - nucleoside diphosphate kinase
The purine salvage pathway • Purine bases created by degradation of RNA or DNA and intermediate of purine synthesis were costly for the cell to make, so there are pathways to recover these bases in the form of nucleotides • Two phosphoribosyl transferases are involved: • APRT (adenine phosphoribosyl transferase) for adenine • HGPRT (hypoxanthine guanine phosphoribosyl transferase) for guanine or hypoxanthine
Salvage of purines Adenine phosphoribosyltransferase (APRT)
Salvage is needed to maintain the purine pool (biosynthesis is not completely adequate, especially in neural tissue) Hypoxanthine-guanine phosphoribosyltransferase (HGPRT) Hypoxanthine + PRPP IMP + Ppi Guanine + PRPP GMP + Ppi Lack of HGPRT leads to Lesch-Nyhan syndrome. Lack of enzyme leads to overproduction of purines which are metabolized to uric acid, which damages cells Salvage of purines
Lesch-Nyhan syndrome • there is a defect or lack in the HGPRT enzyme • the rate of purine synthesis is increased about 200X • uric acid level rises and there is gout • in addition there are mental aberrations • patients will self-mutilate by biting lips and fingers off
Next: Part 2 - biosynthesis of pyrimidine nucleotides
Nucleotide metabolism – Part 2(pyrimidine biosynthesis) By Henry Wormser, Ph.D Professor of Medicinal Chemistry
Synthesis of pyrimidine ribonucleotides • shorter pathway than for purines • base is made first, then attached to ribose-P (unlike purine biosynthesis) • only 2 precursors (aspartate and glutamine, plus HCO3-) contribute to the 6-membered ring • requires 6 steps (instead of 11 for purine) • the product is UMP (uridine monophosphate)
Step 1: synthesis of carbamoyl phosphate • Condensation of glutamine, bicarbonate in the presence of ATP • Carbamoyl phosphate synthetase exists in 2 types: CPS-I which is a mitochondrial enzyme and is dedicated to the urea cycle and arginine biosynthesis) and CPS-II, a cytosolic enzyme used here
Step 1: pyrimidine synthesis CPS-II is the major site of regulation in animals: UDP and UTP inhibit the enzyme and ATP and PRPP activate it It is the committed step in animals
Step 2: synthesis of carbamoyl aspartate • enzyme is aspartate transcarbamoylase (ATCase) • catalyzes the condensation of carbamoyl phosphate with aspartate with the release of Pi • ATCase is the major site of regulation in bacteria; it is activated by ATP and inhibited by CTP • carbamoyl phosphate is an “activated” compound, so no energy input is needed at this step
Step 3: ring closure to form dihydroorotate • enzyme: dihydroorotase • forms a pyrimidine from carbamoyl aspartate • water is released in this process