Enhanced Protein Parsimony Method for Peptide Identification Sensitivity Improvement
60 likes | 153 Vues
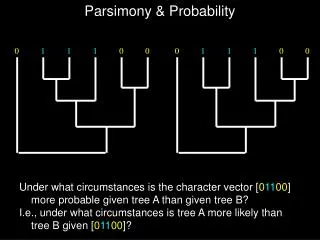
This method selects the smallest set of proteins covering all identified peptides using a sensible principle to avoid bad consequences for FDR filtered identifications. By maximizing covered peptides and considering unique peptides per protein, it significantly improves identification sensitivity.

Enhanced Protein Parsimony Method for Peptide Identification Sensitivity Improvement
E N D
Presentation Transcript
Generalized Protein Parsimony Nathan Edwards Department of Biochemistry and Molecular & Cellular Biology Georgetown University Medical Center
Traditional Protein Parsimony • Select the smallest set of proteins that cover all identified peptides. • Sensible principle, implies • Eliminate equivalent/subset proteins • Unique peptides force proteins into solution • Bad consequences for FDR filtered ids • Impact most tools, even probability based ones
Must ignore some PSMs • Improving peptide identification sensitivitymakes things worse! • False PSMs don't cluster PSMs PSMs 2x Proteins 10%
Must ignore some PSMs • A single additionalpeptideshould not force proteins into the solution
Generalized Protein Parsimony • Weight peptides by number of PSMs • Constrainunique peptides per protein • Maximize covered peptides (PSMs) • Can match filtering FDR to uncovered PSMs • Readily solved by branch-and-bound • Reduces to traditional protein parsimony
Match FDR to uncovered PSMs Traditional Parsimony at 1% FDR: 1085 (609 2+-Unique) Proteins