beta
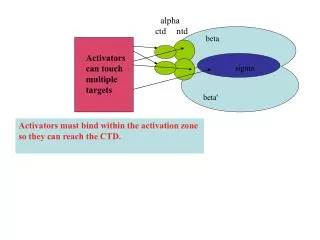
alpha. ctd. ntd. beta. Activators can touch multiple targets. sigma. beta'. Activators must bind within the activation zone so they can reach the CTD. In general, activated promoters typically have a poor enough match

beta
E N D
Presentation Transcript
alpha ctd ntd beta Activators can touch multiple targets sigma beta' Activators must bind within the activation zone so they can reach the CTD.
In general, activated promoters typically have a poor enough match to consensus that polymerase won’t bind. Sometimes the bound RNA polymerase needs activator to change to its active form (details still lacking?) Therefore transcription is dependent on activator spacer Polymerase alpha subunits Sigma r4 Polymerase in active form Sigma r2 They bind here so they can touch polymerase And repressors bind here so they can block polymerase binding (anywhere the polymerase footprints) OPERATORS
Promoter recognition pol Pol bound is a way that may isomerize slowly "binding" (Kb) act closed Activator or other elements Can stabilize intermediate, closed "isomerization" (K2 or Kf) Fully isomerized enzyme and DNA open Only the open complex is resistant to heparin
Conserved promoter elements can also assist RNAP at either the binding or isomerization steps Fenton - PNAS 2001 | vol. 98 | 9020-902 Band shift experiments with each base in the -10 element changed Conclusion: base changes can affect “binding” of duplex DNA up to 10-fold
Because this element is melted in the open complex are the bases also important in single strand form? (This fork junction probe was used to determine the pol-DNA stucture. It was known that the non-template strand was critical for sigma action) Conclusion: bases are of lesser importance in “binding” ssDNA
But when the same exp. is done with heparin the bases are again very important. Conclusion: the element is needed first for pol “binding.” After pol melts the DNA it is important again for isomerization. So both DNA elements and activators can assist at either the binding or pol somerization steps.
free spacer activator -35 -10 binding up closed alpha isomerization open binds AT-rich up element So, pol function is altered by all DNA elements and activators recruiting it and then changing it to the active form before it dissociates to try again.
How to “down-regulate” signaling to the promoter? De-activation of the signalling pathway - whole regulon can be affected. Repression can down-regulate sub-sets of genes. Now consider regulation that targets the pol subunits directly. These work in a way that only a sub-set of genes may be affected. Still not clear how promoter-specificity is built-in in some cases.
Anti-sigmas activator -35 -10 Anti-alphas? alpha
oxidation stress Activates trxA, B and other genes Spx Spx- disulfide Represses srf, hmp and other genes Properties of the dual redox regulator spx (B. subtillis) Spx is not obviously a DNA-binding protein. How does Spx work? Clue from genetics. The cxs-1 mutant in the alpha subunit of RNAP cannot support either activation or repression by Spx.
Activation - trxA, B RNA forms only when: spx is present spx is oxidized (no dtt) the alpha subunit is wt (not cxs1) Nakano… Zuber (2005) Mol. Micro 55:498-510 Interpretation: Oxidized spx activates via the RNAP alpha subunit.
Binding of the 2 together No binding of RNAP alone Now footprint to see what’s bound to the promoter. No binding of oxidized Spx
Interpretation: Oxidized Spx drives RNA polymerase to the promoter even though it is not a conventional DNA-binding protein. Genetics suggest it may do so by triggering a changed state of RNAP by interacting with its alpha subunit. What provides the promoter specificity? Not clear at this point.
Spx repression: it blocks this signal transduction pathway. Nakano ..Zuber (2003) PNAS| vol. 100 | no. 7 | 4233-4238 Mutations in Spx fail to properly down-regulate certain stress-related genes. 2 of these involve “2 component” activators: Activator comA-dependent srf transcription. Activator resD- dependent hmp transcription signal Kinase component Question: how can Spx block these pathways? Phosphorylate activators comA/resD comA/resD bind promoter comA/resD touch alpha comA/resD activate transcription of srf/hmp
Two mutants are useful in repression studies The cxs-1 form of alpha: The RNAP cannot be repressed (or activated - see prior). The cxs-16 form of Spx also cannot do repression. Experiments on next slides. Evidence for above. Use mutants in vitro to find how it blocks comA or resD activators from working
Transcription in vitro Its mutant does not Wt Spx inhibits Wt Spx does not inhibit with alpha mutant ComA transcription of srf The same results are obtained with resD transcription of hmp. Interpretation: Spx interferes with comA/resD -dependent transcription. It requires the wild type alpha subunit for this repression.
Spx binds directly to the polymerase alpha subunit Pull-down exp: His6 tagged Spx His6 tagged Spx bead Ni Ni bead Ni Ni Ni Ni + His6 tagged Spx His6 tagged Spx Ni Ni add alpha and wash Elute with imidazole and run on gel Spx and alpha bind (note controls on this)
Does Spx compete with the comA for alpha? Band shift experiments to see composition of transcription complex. Spx can remove comA from the transcription complex. (note RNAP still there) The Spx mutant cannot The alpha mutant prevents removal comA dependent promoter somewhat similar at resD Interpretation: Spx removes the requiredactivators from the complexes. It appears to act as an anti-alpha.
Interpretation: Spx can act as an anti-alpha, blocking a subset of activator contacts and releasing those activators -35 comA etc -10 Spx alpha Note: somehow Spx activates at other promoters, an unsolved issue.
Next, we will discuss anti-sigmas. But first, a look at the structure of sigma70 in more detail: how it fits into the polymerase core and binds DNA. all from Darst lab - see Bacterial RNA polymerases: the wholo story. Curr Opin Struct Biol. 2003 Feb;13(1):31-9.
Structure of purified whole sigma is inaccessible - here are some fragments of sigma70 (thermophilic) Fragments missing region 1 but with conserved domains 2, 3 and 4. Next see how this interacts with-35 region of DNA
Nothing unusual - primarily a recognition helix in the DNA major groove.
Another view of sigma (when part of the holoenzyme) region 4 region 2
You’ve seen how sigma70 sits on the core before. -35 -10 Flap and coiled-coil are prime determinants on core (underneath the arrows) that bind sigma tightly.
Review of the opening step. open closed
And a closer view of how the DNA strands enter the active site.
Now to the anti-sigma stories - the first concerns sigma28 Sigma28 is needed for the assembly of the bacterial flagellum. Its part of a developmental pathway, in which proteins are provided “just-in-time” (as are needed for assembly). Sigma28 is produced early, but only needed after FlgM protein has been incorporated into the flagellum. Then sigma28-dependent proteins are produced and complete the assembly. Question: how is sigma 28 activated after FlgM is depleted?
Short answer: FlgM is an anti-sigma. It binds and inactivates sigma28 before going into the flagellum. The depletion of FlgM releases sigma28 and thus activates transcription . Next data from Chadsey et al(1998) Genes Dev. 12:3123-36
Basic properties of the transcription system. sigma28 activates the fliC promoter but not a sigma70 promoter flgM inhibits fliC transcription
Lane 2 shows that flgM binds sigma28 in a GST-pull down assay
FlgM works by blocking promoter binding by sigma28 holoenzyme E = core
The next slides compare the structure of sigma28 in its FlgM complex to that of free sigma 70 (28 is homologous to 70 in regions they share). Sorenson MK, Ray SS, Darst SA.Crystal structure of the flagellar sigma/anti-sigma complex sigma(28)/FlgM reveals an intact sigma factor in an inactive conformation. Mol Cell. 2004 Apr 9;14(1):127-38 Likely interpretation: flgM binds sigma28 and prevents its use in promoter recognition: An anti-sigma
Sigma28 is condensed when bound to the anti-sigma FlgM Interpretation: the 2 regions should not be able to bind DNA simultaneously region 4 region 4 region 2 region 2
This shows that flgM binds sigma28 in a way that competes with binding the coiled-coil site on core polymerase. So, it inhibits core binding. (flap binding also inhibited - not shown here).
This is how sigma28 would need to unfold in order to bind core polymerase and function. Because sigma70 requires unknown conformational changes to bind core, this may be a model for that. Condensed (by anti-sigma) Structure in polymerase One more example of an anti-sigma
Stress in the periplasm leads to degradation of ResE Now sigmaE can function sigmaE holoenzyme transcribes proteins in response to stresses that damage proteins in the bacterial periplasmic space. outside RseA is an anti-sigma and blocks sigmaE periplasm RseA cytoplasm sigmaE
The structure of a sigmaE fragment with the anti-sigma RseA. Note that it also condenses a sigma such that the DNA-binding regions cannot bind -10/-35 simultaneously. Sigma r2 Rse Sigma r4 Campbell EA, Mol Cell. 2003 Apr;11(4):1067-78
A closer look confirms that it forms a surface that can condense sigma regions 2 and 4
So, inhibition by anti-sigmas and anti-alphas can block transcription of targeted set of genes. homework - use data from the lecture to explain what interactions might be made defective by the cxs-1 and cxs-16 mutations.