1 / 16
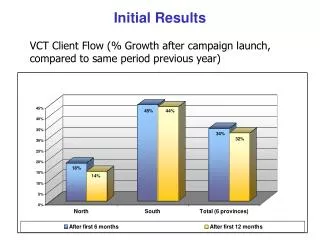
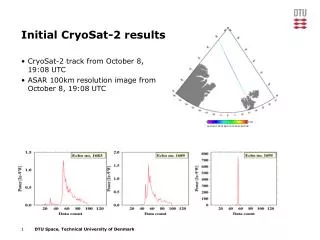
HSC RNAseq Initial Results
160 likes | 266 Vues
Quality control analysis of RNA-seq results from Ariel Paulson, focusing on read trimming, alignment using Bowtie and Tophat, and unique read alignments. Segmented at 35bp segments, allowing only UM reads with 2 mismatches each.
Télécharger la présentation 

HSC RNAseq Initial Results
An Image/Link below is provided (as is) to download presentation
Download Policy: Content on the Website is provided to you AS IS for your information and personal use and may not be sold / licensed / shared on other websites without getting consent from its author.
Content is provided to you AS IS for your information and personal use only.
Download presentation by click this link.
While downloading, if for some reason you are not able to download a presentation, the publisher may have deleted the file from their server.
During download, if you can't get a presentation, the file might be deleted by the publisher.
E N D
Presentation Transcript
HSC RNAseqInitial Results Ariel Paulson
Data QA/QC \\dm3\solexa-analysis\Li_Linheng\ave\Li_Linheng-2011-07-25\BB027YABXX\fastqc\fastqc_plots.htm Col2_3GFP Mega5FU READS TRIMMED TO 70bp long
QC FAIL NO MATCH TRUE NO MATCH MULTI ALIGN UNIQUE ALIGN UNIQUE AT 0MM TAGS % READS
Alignment using Bowtie 0.12.7 + Tophat, 1.3.1, only allow for UM reads. Broken into 2 segments @ 35bp each, 2mm each
More Related