3 10
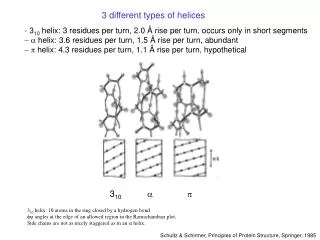
3 different types of helices. - 3 10 helix: 3 residues per turn, 2.0 Å rise per turn, occurs only in short segments - a helix: 3.6 residues per turn, 1.5 Å rise per turn, abundant p helix: 4.3 residues per turn, 1.1 Å rise per turn, hypothetical. 3 10. a. p.

3 10
E N D
Presentation Transcript
3 different types of helices • - 310 helix: 3 residues per turn, 2.0 Å rise per turn, occurs only in short segments • - a helix: 3.6 residues per turn, 1.5 Å rise per turn, abundant • p helix: 4.3 residues per turn, 1.1 Å rise per turn, hypothetical 310 a p 310 helix: 10 atoms in the ring closed by a hydrogen bond. fy angles at the edge of an allowed region in the Ramachandran plot. Side chains are not as nicely staggered as in an a helix. Schultz & Schirmer, Principles of Protein Structure, Springer, 1985
Leucine zippers Leucine zippers are heptad repeats containing a leucine at position 4 with almost complete conservation. Example: transcription factor GCN4 Leucine zippers form a-helices with a helical repeat of 3.5 residues per turn (instead of 3.6 residues per turn as in a conventional a-helix). Plotted on a helical wheel, the leucine residues all face the same side of the helix. Residues in positions a and d are hydrophobic or, at least, uncharged. Carl Branden & John Tooze, Introduction to Protein Structure, Garland, 1998
Leucine zippers form two parallel coiled-coil a helices, where the hydrophobic side chains in positions a and d of the heptad repeats form a hydrophobic core between the helices with the leucine residues facing each other. The side chains immediately outside this core (positions e and g) are frequently charged and can either promote or prevent dimer formation. The GCN4 basic region-leucine zipper binds DNA as a dimer of two uninterrupted a helices. Leucine zipper homodimers and heterodimers can recognize different DNA sequences, as indicated by the red and blue regions of the DNA.
Additional Domain swapping antigen binding site Fab fragment of conventional antibody antibody binding sites with carbohydrates from HIV gp120 new conventional conventional Domain swapping generates a dimer with increased binding surface to multiple carbohydrate chains. Furthermore, additional new binding sites are created, enhancing the binding affinity. > 40 cases of domain swapping identified. Some of them are crystallization artifacts, while some of them are physiologically relevant. Science 300, 2065 (2003) Prot. Sci. 11, 1285 (2002)
What is a domain? • a segment of similar amino acid residues identified in different proteins (by sequence alignment) • an independent folding unit • Domains are often arranged like beads on a string, connected by flexible linkers. Example: domain architectures of apoptotic proteins Science, 291, 1279 (2001) Compare, however, with PEST domains! (PEST: domains rich in Pro, Glu, Ser and Thr, which enhance the degradation rate of proteins by the proteasome) PEST domains are unstructured.
Protein function derived from genome comparison Some pairs of interacting proteins have homologs in another organism fused into a single protein chain. Searchingsequences from many genomes revealed 6809 such putative protein-protein interactions in Escherichia coli and 45,502 in yeast. Two homologous enzymes are fused in human. Two nonsequential enzymes from the histidinebiosynthesis pathway are fused in yeast. data base of interacting proteins: http://dip.doe-mbi.ucla.edu/ Science 285, 751 (1999)
Amino acid composition of unstructured versus structured proteins Amino acid frequencies in%. Asterisks identify amino acids that are at least two times more or less frequent than in an average globular protein in the Protein Data Bank. Trends in Biochem. Sci. 27, 527 (2002)
Not all proteins have a defined 3D structure Up to 50% of eukaryotic proteinshave at least one long (>50 residues) disordered region. 11% of proteins inSwiss-Prot are probably fully disordered These proteins are likely to assume a defined structure in complex with other proteins. Two proteins can be unstructured by themselves, but can form a specific complex of defined globular structure, when together. p27Kip1 complexed with cyclin-dependent kinase 2 (Cdk2) and cyclin A [CycA; ProteinData Bank (PDB) number 1JSU] Nature 415, 549 (2002) Trends in Biochem. Sci. 27, 527 (2002)
Protein folding-unfolding equilibria and denaturation Folded protein structures are in equilibrium with partially or fully unfolded forms. The conformational stability equals the free energy change DG of the unfolding reaction: F U DG is small: 5-15 kcal/mol for proteins with 70-200 residues. For comparison: DG = 4-5 kcal/mol for the formation of a single H-bond between two water molecules. • Unfolded polypeptide chains assume a ‘random coil conformation’. • Unfolding can be achieved by • increased temperature (= heat denaturation) • decreased temperature (= cold denaturation) • chemicals (e.g. 6 M urea or guanidinium chloride) urea guanidinium ion Urea and guanidinium chloride denature proteins because they can form more stable H-bonds to the backbone amides than water.
Circular Dichroism (CD) Spectroscopy Circular Dichroism is observed when optically active matter absorbs left and right hand circular polarized light slightly differently. Linear polarized light can be viewed as a superposition of opposite circular polarized light of equal amplitude and phase. When this light passes through a sample with a different absorbance A for the two components, the amplitude of the stronger absorbed component will smaller than that of the less absorbed component. The consequence is that a projection of the resulting amplitude now yields an ellipse instead of the usual line. The CD signal at 222 nm is often used to monitor folding-unfolding equilibria as a function of temperature. http://www-structure.llnl.gov/cd/cdtutorial.htm
Folding-Unfolding Equilibria by CD and Fluorescence unfolded folded The sigmoidal shape of the curve is a consequence of the cooperativity of unfolding. Fluorescence mostly relies on the fluorescence of tryptophan residues which is different for solvated versus buried side chains.
Amyloid Fibres Many proteins can become insoluble by the formation of amyloid fibres. These fibres consist of 4x2 sets of infinitely long parallel b-sheets stacked in two layers. Cryoelectron microscopy image Model of a fibril http://people.cryst.bbk.ac.uk/~ubcg16z/amyloid/amyloid.html
Chaperones help proteins fold GroEL and GroES are E. coli proteins. GroEL forms a cavity, where unfolded protein (I) can bind. ATP is required to form the complex with GroES. The release of the folded (N) or still unfolded protein requires again ATP. GroEL acts by partial unfolding of misfolded proteins, presumably by stretching the polypeptide chain in the cycle between open and closed conformation of the GroEL ring. mcdb.colorado.edu/courses/ 3280/class03-3.htm Science 284, 823 (1999)l
cis-trans isomerization of peptide bonds preceding Pro cis trans In the peptide Ac-Ala-X-Pro-Ala-Lys-NH2, the amount of cis-peptide bond varies between 6% for X=Pro and 38% for X=Trp. (Nat. Struct. Biol. 6, 910 (1999)) Despite the relative rigidity of the side chain of proline, frequent occurance of prolines reduces the chances for globular structures due to the cis-trans heterogeneity.
Many heat shock proteins are chaperones. Proline-cis-trans isomerases can be considered as chaperones, as the cis-trans isomerization step is slow compared with bond rotation around single bonds. Proteins with more than one disulfide bond often form non-native disulfide bonds during folding which must be corrected. DsbA and DsbC in E. coli contain thioredoxin-like domains which assist in reduction and oxidation of disulfide bonds. DsbA contains two cysteines separated by two residues. The disulfide bond between these two cysteines is very reactive and act as an oxidizing agent for other cysteines. DsbA
Enzymes can provide a non-aqueous environment arachidonic acid cyclooxygenase PGG2
Enzymes function by stabilizing the transition state Example: catalytic antibodies. The antigen-binding sites are hypervariable. Antibodies can be selected which bind to any molecular target of interest.
An antibody raised against a transition state analog catalyzes the reaction from chorismate to prephenate: Note: catalytic antibodies occur naturally, e.g. with proteolytic function in autoimmune diseases. Hilvert lab, ETH-Zűrich
The Role of Water in Intermolecular Recognition NMR experiments and molecular simulations have shown that the residence times of buried hydration water molecules is shorter than 1 ms. On protein and DNA surfaces, the residence times are shorter than 1 ns. Solvation-desolvation processes present no kinetic impediment to protein function. Science 254, 974 (1991) J. Mol. Biol. 282, 859 (1998)
But: water molecules can play a crucial structural role! Example: in the E. coli Trp-repressor/DNA complex, the specificity determining contacts between the protein and the DNA bases are mediated by water molecules. W denotes water molecules. Base-pairs on the left, protein on the right. The repressor binds with a helix-turn-helix motif to the major groove of the DNA. The Trp-repressor does not recognize ‘hydrated DNA’. It is the energetically favourable arrangement of the complex which determines the importance of the interfacial water molecules in the protein-DNA complex. Nature 335, 321 (1988)