Advanced Bioinformatics
The Advanced Bioinformatics Group is embarking on an exploratory project to analyze gene expression in yeast. Our objectives include creating a top 100 list of highly expressed genes and investigating correlations between gene expression, transcript attributes, and genetic metrics such as GC content, transcript length, intron length, and codon usage. Utilizing tools like Tophat and Cufflinks, we aim to visualize interaction networks and establish informative Perl scripts for comprehensive data analysis. This project represents both an end and a new beginning in our research journey.

Advanced Bioinformatics
E N D
Presentation Transcript
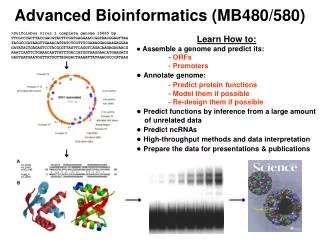
Advanced Bioinformatics Group Medicago Basic Project: Gene expression in yeast
Members • JenteOttenburghs • Lifei Li • Yuebang Yin • Nick Brouwers
Objectives • Make a top 100 of the most highly expressed genes • Find correlations between gene expression & transcripts • GC content • Transcript length • Intron length • Codon usage • (optional) Visualization of an interaction network Make use of Tophat, Cufflinks, perl etc.
Steps • Run Tophat • Use Tophat output as Cufflinks input • Use Cufflinks output to build Perl scripts • Show our analysis and hopefully visualization
Plan: Data gathering • Create output with Cufflinks • Build Fasta from transcripts.gtf and genome.fa(all) • Vizualize interaction networks (Yuebang) • Build scripts to: • analyse our Fasta, GC content, length etc. (Nick) • implement gene expression (Lifei Nick) • sort gene expression (Lifei and Nick) • show intron length (all together) • determine condonusage (Jente)