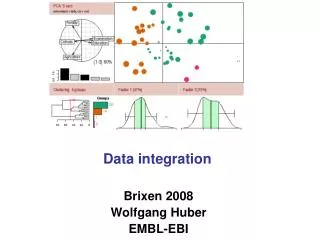
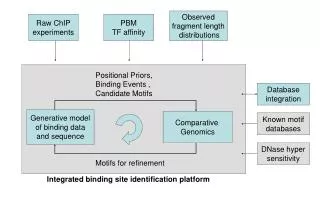
EM Database Data integration
The EMDB provides a robust platform for integrating and coordinating protein crystallography software through the CCP4 interface. It offers validated tools based on bond geometry and free R-factor, alongside a complete repository that features user-friendly visualization and sophisticated searching capabilities. Both novices and experts benefit from a well-designed interface with extensive validation information, including links to PDB and experimental data. Desirable features include advanced map manipulation tools and integration with cell structure data for enhanced structural analysis.


EM Database Data integration
E N D
Presentation Transcript
EM Database Data integration
Role model – protein crystallography CCP4 – common interface to coordinate and integrate most of the software in the field; workshops, training; well established validation tools based on bond geometry, free R factor PDB – Complete public repository with user-friendly visualization and integration tools, sophisticated searching, validation information, used by experts and non-specialists
Basic featuresfor the EMDB • A nice front page – for both novice and expert users • Searching on entry number, author, keyword,.. • A friendly data page with a picture of the map • Bug-free map header • Links to sequence and function databases • Experimental information • Validation information • Links to and from the PDB
Desirable featuresfor the EMDB • A Rasmol for maps • Tools for moving and aligning maps • A button to shift the EM map from the center of the box to the origin of the coordinate system, needed for symmetry operations and for matching up with pdb files • Searching densities by cross correlation • Integrate with cell structure data • Validation with Rosenthal/Henderson tilt data • Fold searching for maps at <7 Å • Full integration into the PDB