Chapter 9 The cell nucleus and chromosome
1.03k likes | 1.58k Vues
Chapter 9 The cell nucleus and chromosome. Overall structure. 9.1 Structure of nuclear envelope 9.2 Chromatin DNA second structure DNA first structure complexity Chromatin protein Basic unit of chromatin-----nucleosomes Chromatin assemble Chromatin type

Chapter 9 The cell nucleus and chromosome
E N D
Presentation Transcript
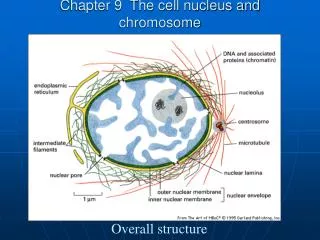
Chapter 9 The cell nucleus and chromosome Overall structure
9.1 Structure of nuclear envelope • 9.2 Chromatin • DNA second structure • DNA first structure complexity • Chromatin protein • Basic unit of chromatin-----nucleosomes • Chromatin assemble • Chromatin type • Chromatin and regulation • 9.3 Chromosome • 9.4 The nucleolus
Internal structure • Nuclear envelope • Nuclear pores • Nucleolus • Chromatin • Nuclear lamina (nuclear cortex) • Nuclear matrix (internal fibrils)
9.1 Structure of nuclear envelope Nucleus Membrane 9.1.1Nuclear envelope Nuclear pores complex
outer membrane (7.5nm) connected to ER 9.1.1 Nuclear membrane Nuclear pore Peri-nuclear space(20-40nm) Inner membrane (7.5nm) contact with nucleus lamina
9.1. 2 The nuclear pores 0.1um Side view of nuclear pore
Size and density of nuclear pore in different mammalian cells
Nuclear pore density of numbers of the nuclear pores per nucleus
First observation of nuclear pore in 1950 by A. Callan and S. Tomlin, named as nuclear pore complex (NPC) in 1959 by Watson • Pore size is variable according to the cell types and tissue • In general 40 - 100 nm • Numbers of pores in given area • low in cells with slow metabolism and at times of low activity during cell cycle • high in cells after cell division and with higher activity of RNA transport and protein synthesis
Cytoplasmic ring structure inner diameter of 90 nm and outer diameter of 120 mm • eight small subunits in octagonal arrangement • sometimes, in 16 or 24 subunits • Spokes, link cytoplasmic ring structure with nuclear ring • Central plug, 35 nm in diameter, transporter • Nuclear ring: nuclear basket or fish-trap small molecules arranged along with the ring structure in the cytoplasmic side
Nuclear pore complex component Elaborate structure of approx. 1000 proteins forming, protein lined aqueous channel approx 9nm diameter Protein fibrils protrude each side of complex - form cage-like structure on nuclear side Typical cell contains 3000-4000 pore complexes Each pore, on average, imports 100 histone molecules per minute and exports 6 small ribosomal subunits Gp120 and p62
9.1.3.1 Nuclear protein transport mechanism basic definition ◆nuclear protein ◆nuclear localization signals ,NLS ◆nuclear export signals, NES ◆importin ◆exportin
9.1.3.2 Mechanism of export from nucleus Most of traffic moving out of nucleus consists of different types of RNA molecules RNA moves through nuclear pore as complex of ribonucleoprotein (RNP) Protein component of RNP contains a nuclear export signal (NES) that is recognised by export proteins mRNA is bound by hnRNP only after fully spliced so only mature RNA is exported
Nuclear RNA and protein export ◆mRNA export ●异质核糖核蛋白 (heterogeneous ribonucleoprotein, hnRNP) ●信使RNP(messenger RNP, mRNP) ●mRNA的输出 ◆nuclear protein ◆snRNA export
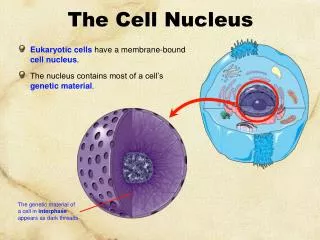
9.2 chromatin • Contains DNA and protein
9.2.2 DNA first structure complexity • Three classes of DNA • 1、unique repeat DNA • code for genes and unique material • 2.6% of unique genome is transcribed in frogs • interspersed repeated (moderately repetitive) • 2、tandemly repeated (highly repetitive)
3、Moderately repetitive Coding • 40-50% of genome is this type • Coding • code for genes needed in large amounts • rDNA to make rRNA (has 250 copies per human genome) Noncoding
Noncoding • up to 300 bp in length (ALU sequences) • 10,000 - 100,000 copies per genome • often interspersed between unique sequences • may be mobile (jumping genes Short interspersed elements, SINEs Eg. ALU sequence(<500bp) Noncoding Long interspersed elements, LINEs (>1000bp)
在基因组的重复序列中有一个大家族,被称为Alu家族。每一个Alu片段大约有300个核苷酸。在人类基因组中,大约有140万个Alu序列,占整个基因组的10%。在基因组的重复序列中有一个大家族,被称为Alu家族。每一个Alu片段大约有300个核苷酸。在人类基因组中,大约有140万个Alu序列,占整个基因组的10%。 • Alu片段在过去的3000万年间快速而大量地富集于基因组GC碱基含量高的区域内,比GC贫乏区的Alu片段含量高13倍。有趣的是,GC富集的区域内基因的密度也比GC贫乏区大。显然,Alu序列与基因在基因组中的进化可能有某种相关性。 • 2003年5月,以色列科学家在美国《科学》周刊上发表了他们的研究结果,揭示出Alu片段在基因剪接(splicing)过程中插入到完整的mRNA中的分子机理,表明重复序列在进化过程中可以用来帮助形成新基因。
Minisatellite DNA(12-100bp,3000 copy) Highly repetitive Microsatellite DNA(1-5bp form 5-100bp) Satellite DNA(5-100bp) • 10-15% of genome (mammals) • short stretches (as little as a dozen basepairs) • GC content may be different than genome average • location - only at a few locations; centromeres • often repeated up to 100-100,000 times in tandem • role is unknown - speculation is they are involved in the mechanics of meiosis
Nucleosome protein H2B,H2A,H3,H4, conservative Histone protein Sructure protein H1,link function No-histone protein : sequence specific DNA binding protein 9.2.3 Chromatin protein Chromatin protein
No-histone protein • No-histone protein diversity, heterogeneous Nonhistone chromosomal proteins include a large number of widely diverse structural, enzymatic, and regulatory proteins. • Specific recognition of DNA structure Helix –turn –helix motif Zinc- finger-motif Leucine-zipper motif Helix-loop-helix motif HMG ( high mobility group proteins)-box motif • Multifunction
The helix-turn-helix motif ●α-helix form homeotypic dimer ●α-helix dimer link with rotational peptide ●recognition helix ●orther helix no specific
zinc finger motif Cys2/His2 and Cys2/Cys2
The leucine zipper motif The leucine zipper contains four or five leucine residues spaced at intervals of seven amino acids, resulting in their hydrophobic side chains being exposed at one side of a helical region. This region serves as the dimerization domain for the two protein subunits, which are held together by hydrophobic interactionsbetween the leucine side chains.
Helix-loop-helix (HLH) motif This motif is characterized by two -helical segments separated by an intervening loop. 它与螺旋-转角-螺旋结构的差别在于∶两个螺旋的一侧还有一段疏水链,这样当螺旋-环-螺旋结构位于两个多肽之间时,这两个疏水的侧链就会将两个多肽链连在一起形成类似亮氨酸拉链的结构
B Histone ◆The histones are small proteins containing a high proportion of basic amino acids (arginine and lysine) that facilitate binding to the negatively charged DNA molecule. ◆There are five major types of histones--called H1, H2A, H2B, H3, and H4 . which are very similar among different species of eukaryotes , very conservative ◆Ratio of Histone and DNA is 1:1
Nucleosome protein H2B,H2A,H3,H4, conservative Histone protein Sructure protein H1,link function No-histone protein sequence specific DNA
Nucleosome Nucleosome is basic structural unit of chromatin。 ◆200 bp DNA and four type histone; ◆H2A、H2B、H3、H4 ,every type two copy form octameric core ,is key structure of Nucleosome ◆ 146bp DNAsurround octameric core 1.75 circle ,H1 play stablization function in chromatin formation ◆ Neighborhoodnucleosome connect with linker DNA link (about ~90bp)
9.2.6 Chromatin type • Euchromatin: open structure – active genes • Heterochromatin: highly packaged structure – inactive genes
◆常染色质(euchromatin) ◆异染色质(heterochromatin) ●结构性异染色质(active chromatin) ●兼性异染色质(inactive chromatin)
9.2.7 Chromatin and regulation • Chracterristic of active chromatin • Nucleosome and gene regulation