Chapter 13 Enzyme Kinetics
Chapter 13 Enzyme Kinetics. Outline. What characteristic features define enzymes ? Can the rate of an enzyme-catalyzed reaction be defined in a mathematical way ? What equations define the kinetics of enzyme-catalyzed reactions ? What can be learned from the inhibition of enzyme activity ?


Chapter 13 Enzyme Kinetics
E N D
Presentation Transcript
Outline • What characteristic features define enzymes ? • Can the rate of an enzyme-catalyzed reaction be defined in a mathematical way ? • What equations define the kinetics of enzyme-catalyzed reactions ? • What can be learned from the inhibition of enzyme activity ? • What is the kinetic behavior of enzymes catalyzing bimolecular reactions ? • How can enzymes be so specific ? • Are all enzymes proteins ? • Is it possible to design an enzyme to catalyze any desired reaction ?
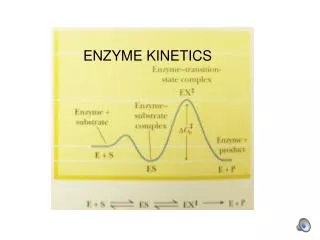
Virtually All Reactions in Cells Are Mediated by Enzymes • Living systems use enzymes to accelerate and control the rates of vitally important biochemical reactions (see Figure 13.1). • Enzymes provide cells with the ability to exert kinetic control over thermodynamic potentiality. (glucose pyruvate) • Enzymes are the agents of metabolic function. • Most enzymes are proteins. • Some enzymes require cofactors or coenzymes.
Virtually All Reactions in Cells Are Mediated by Enzymes Figure 13.1 Reaction profile showing the large free energy of activation for glucose oxidation. Enzymes lower ΔG‡, thereby accelerating rate.
13.1 What Characteristic Features Define Enzymes ? • Catalytic power is defined as the ratio of the enzyme-catalyzed rate of a reaction to the uncatalyzed rate. • Specificity is the term used to define the selectivity of enzymes for their substrates. • Regulation of enzyme activity ensures that the rate of metabolic reactions is appropriate to cellular requirements. • Coenzymes and cofactors are nonprotein components essential to enzyme activity.
13.1 What Characteristic Features Define Enzymes ? • Enzymes can accelerate reactions as much as 1016 over uncatalyzed rates. • Urease is a good example: • Catalyzed rate Uncatalyzed rate Ratio 3x104/sec 3x10-10/sec 1x1014 • Enzymes selectively recognize proper substrates over other molecules. • Enzymes produce products in very high yields - often much greater than 95%. • Specificity is controlled by structure - the unique fit of substrate with enzyme controls the selectivity for substrate and the product yield.
Enzymes are the Agents of Metabolic Function Figure 13.2 The breakdown of glucose by glycolysis provides a prime example of a metabolic pathway with many sequential steps. All pathways are regulated but not all enzymes are regulated . Regulation of glycolysis occurs at several points, the major control point is through the enzyme phosphofructokinase I.
90% yield in each step = 35% over 10 steps Figure 13.3 Yields in biological reactions must be substantially greater than 90%.
Enzymes Nomenclature As in organic, there is a system of nomenclature devised for enzymes as well as a lot of commonly used names. There are six main classes used for naming enzymes. Know these six classes. There is also a numerical assignment for each enzyme developed by the Enzyme Commission. E.g.: EC 1.1.1.1 denotes alcohol dehydrogenase Its systematic name is: alcohol:NAD+ oxidoreductase donor:acceptor main class
The Six Classes of Enzymes Class# Class and reaction catalyzed 1.Oxidoreductases (dehydrogenases) oxidation-reduction reactions 2.Transferases group transfer reactions 3. Hydrolases hydrolysis reactions 4. Lyases lysis, forming a double bond 5. Isomerases isomerization reactions 6.Ligases (synthetases) joining of two substrates, uses ATP
Enzyme Nomenclature Provides a Systematic Way of Naming Metabolic Reactions Know the six main classes.
Coenzymes and Cofactors Are Nonprotein Components Essential to Enzyme Activity Coenzymes are organic cofactors.
13.2 A Mathematical statement of the Rate of an Enzyme-Catalyzed Reaction • Kinetics is the branch of science concerned with the rates of reactions. • Enzyme kinetics seeks to determine the maximum reaction velocity that enzymes can attain and binding affinities for substrates and inhibitors. • Analysis of enzyme rates yields insights into enzyme mechanisms and metabolic pathways. • This information can be exploited to control and manipulate the course of metabolic events.
Several kinetics terms to understand • rate or velocity of reaction • rate constant • rate law • order of a reaction • molecularity of a reaction
Chemical Kinetics Provides a Foundation for Exploring Enzyme Kinetics • Consider a reaction of overall stoichiometry as shown: • The rate is proportional to the concentration of A
Chemical Kinetics Provides a Foundation for Exploring Enzyme Kinetics • The simple elementary reaction of A→P is a first-order reaction, v = k[A] and exponent on [A] is 1. • Figure 13.4 shows the course of a first-order reaction as a function of time. • This is a unimolecular reaction. • For a second order reaction, the rate law is: • v = k[A][B]. This reaction may be bimolecular or unimolecular depending on the mechanism. • Kinetics cannot prove a reaction mechanism. • Kinetics can only rule out various alternative hypotheses because they don’t fit the data.
The Time-Course of a First-Order Reaction Figure 13.4 Plot of the course of a first-order reaction. The half-time, t1/2 is the time for one-half of the starting amount of A to disappear.
Catalysts Lower the Free Energy of Activation for a Reaction • A typical enzyme-catalyzed reaction must pass through a transition state. • The transition state sits at the apex of the energy profile in the energy diagram. • The reaction rate is proportional to the concentration of reactant molecules with the transition-state energy. • This energy barrier is known as the free energy of activation, ΔG‡. • Decreasing ΔG‡ increases the reaction rate. • The activation energy is related to the rate constant by:
Catalysts Lower the Free Energy of Activation for a Reaction Figure 13.5 Energy diagram for a chemical reaction (A→P) and the effects of (a) raising the temperature from T1 to T2, or (b) adding a catalyst.
The Transition State Understand the difference between G and G‡ • The overall free energy change for a reaction is related to the equilibrium constant. • The free energy of activation for a reaction is related to the rate constant. • It is extremely important to appreciate this distinction.
13.3 What Equations Define the Kinetics of Enzyme-Catalyzed Reactions ? • Simple first-order reactions display a plot of the reaction rate as a function of reactant. concentration that is a straight line (Figure 13.6) • Enzyme-catalyzed reactions are more complicated. • At low concentrations of the enzyme substrate, the rate is proportional to S, as in a first-order reaction. • At higher concentrations of substrate, the enzyme reaction approaches zero-order kinetics. • This behavior is a saturation effect.
13.3 What Equations Define the Kinetics of Enzyme-Catalyzed Reactions ? Figure 13.6 A plot of v versus [A] for the unimolecular chemical reaction, A→P, yields a straight line having a slope equal to k.
As [S] increases, kinetic behavior changes from 1st order to zero-order kinetics Figure 13.7 Substrate saturation curve for an enzyme-catalyzed reaction.
The Michaelis-Menten Equation is the Fundamental Equation of Enzyme Kinetics • Louis Michaelis and Maud Menten's theory. • It assumes the formation of an enzyme-substrate complex. • It assumes that the ES complex is in rapid equilibrium with free enzyme. • Breakdown of ES to form products is assumed to be slower than 1) formation of ES and 2) breakdown of ES to re-form E and S. • Briggs and Haldane later introduced the steady state assumtion.
The Michaelis-Menten Equation is the Fundamental Equation of Enzyme Kinetics k1 k2 E + S ES E + P k-1 E = enzyme concentration. S = Substrate concentration. ES = Enzyme-substrate complex concentration (noncovalent). P = product concentration. k1 = rate constant for formation of ES from E + S. k-1 = rate constant for decomposition of ES to E + S. k2 = rate constant for decomposition of ES to E + P.
k1 k2 E + S ES E + P k-1 Development of the Michaelis-Menton Equation 1. The overall rate of product formation: v = k2 [ES] 2. Rate of formation of [ES]: vf = k1[E][S] 3. Rate of decomposition of [ES]: vd = k-1[ES] + k2 [ES] 4. The steady state assumption requires that: Rate of ES formation = Rate of ES decomposition 5. So: k1[E][S] = k-1[ES] + k2 [ES]
Michaelis-Menton Derivation 6. In solving for [ES], use the enzyme balance to eliminate [E]. ET = [E] + [ES] 7. k1 (ET - [ES])[S] = k-1[ES] + k2 [ES] k1 ET[S] - k1[ES][S] = k-1[ES] + k2 [ES] 8. Rearrange and combine [ES] terms: k1 ET[S] = (k-1 + k2 + k1 [S])[ES] k1 ET[S] 9. Solve for [ES] = ----------------------- (k-1 + k2 + k1 [S])
Michaelis-Menton Derivation ET[S] 10. Divide through by k1:[ES] = ----------------------- (k-1 + k2)/k1 + [S] 11. Defined Michaelis constant: KM = (k-1 + k2) / k1 12. Substitute KM into the equation in step 10. 13. Then substitute [ES] into v = k2 [ES] from step1 and replace Vmax with k2 ET to give: Vmax[S] vo = ----------- KM + [S]
[ES] Remains Constant Through Much of the Enzyme Reaction Time Course Figure 13.8 Time course for a typical enzyme-catalyzed reaction obeying the Michaelis-Menten, Briggs-Haldane models for enzyme kinetics. The early state of the time course is shown in greater magnification in the bottom graph.
The dual nature of the Michaelis-Menten equation Combination of 0-order and 1st-order kinetics • When S is low, the equation for rate is 1st order in S. • When S is high, the equation for rate is 0-order in S. • The Michaelis-Menten equation describes a rectangular hyperbolic dependence of v on S. • The relation of the “rectangular hyperbola” to the enzyme kinetics profile is described in references at the end of the chapter.
Understanding Vmax The theoretical maximal velocity • Vmax is a constant. • Vmax is the theoretical maximal rate of the reaction - but it is NEVER achieved in reality. • To reach Vmax would require that ALL enzyme molecules are tightly bound with substrate. • Vmax is asymptotically approached as substrate is increased.
Understanding Km The "kinetic activator constant" • Km is a constant. • Km is a constant derived from rate constants. • Km is, under true Michaelis-Menten conditions, an estimate of the dissociation constant of E from S. • A measure of ES binding. • Small Km means tight binding; large Km means weak binding. Where k2 is small then Km ≈ Kd.
Table 13.3 gives the Km values for some enzymes and their substrates
Table 13.3 gives the Km values for some enzymes and their substrates
The Turnover Number Defines the Activity of One Enzyme Molecule A measure of catalytic activity • kcat, the turnover number, is the number of substrate molecules converted to product per enzyme molecule per unit of time, when E is saturated with substrate. A measure of rate of enzyme activity. • If the M-M model fits, k2 = kcat = Vmax/Et. • Values of kcat range from less than 1/sec to many millions per sec.
The Turnover Number Defines the Activity of One Enzyme Molecule
The Ratio kcat/Km Defines the Catalytic Efficiency of an Enzyme The catalytic efficiency: kcat/KmAn estimate of "how perfect" the enzyme is • kcat/Km is an apparent second-order rate constant. • It measures how the enzyme performs when S is low. • The upper limit for kcat/Km is the diffusion limit - the rate at which E and S diffuse together. • The maximum rate of diffusion for small molecules is 109 M-1-sec-1.
The Ratio kcat/Km Defines the Catalytic Efficiency of an Enzyme Superoxide dismutase O2¯· (radical) 1 x 106 5 x 10-4 2 x 109 α-Chymotrypsin Acetyl-Phe-amide 1.4 x 10-1 1.5 x 10-2 9.3
Units of Enzyme Activity Terms in discussing enzyme activity Units of enzyme activity: :mol S/min mol S/sec = katal Specific activity: (to follow purification, see p 99) :mol S/min/mg E mol S/sec/kgm E Molecular activity: (turn-over number, TON = kcat ) :mol S/min/:mol E mol S/sec/mol E
Turnover Number Example calculation An enzyme (1.84 :gm, MW 36800), in presence of excess substrate catalyzes at a rate of 4.2 :mol substrate/min. Calculate the TON. (:mol S/min/:mol E) 1.84:gm :mol E: = --------------------- = 5 x 10-5:mol E 36800 :gm/:mol Vmax 4.2 :mol S/min TON = ------ = ----------------------- = 84000 min-1 Et 5 x 10-5:mol E
Michaleis-Mention Equation Example calculation The rate of an enzyme catalyzed reaction is 35:mol/min at [S] = 10-4 M. KM for this substrate is = 2 x 10-5 M. Calculate the rate where [S] = 2 x 10-6 :mol/min. VM [S] VM (10-4) v = ------------- so 35 = --------------------- KM + [S] (2 x 10-5) + (10-4) And VM = 1.2(35) = 42 :mol/min (42)(2 x 10-6) (84 x 10-6) v = -------------------------- = ------------- = 3.8 :mol/min-1 (2 x 10-5) + (2 x 10-6) (22 x 10-6)
Specific Activity • Assume that an assay of 0.8 ml of the crude extract gives 0.518 activity units. • (3800/0.8)(0.518) = 2460 units in the 3.8 l. • 2460 units/22800 mg protein = 0.108 units/mg • As purification proceeds, the specific activity increases due to loss of extraneous protein. • By purifying to a constant specific activity, one has reached a limit in purification. Table 5-1, p. 99
Graphical Determination of KM and VM The Michaleis-Menton plot only permits an estimate of Vmax, so KM is also an estimate at VM/2. There are several graphical methods which provide a better determination of VM and KM. • We will focus on the Lineweaver-Burk equation which is obtained by taking the reciprocal of the Michaleis-Menton equation (See the next slide). • A Lineweaver-Burke plot is frequently referred to as a double reciprocal plot since one plots 1/v vs 1/[S]. • The plot gives a straight line which has a slope of KM/VM, a y-intercept of 1/VM and an x-intercept of -1/KM.
Linear Plots Can Be Derived from the Michaelis-Menten Equation Be able to develop this equation Lineweaver-Burk: Begin with v = Vmax[S]/(Km + [S]) and take the reciprocal of both sides. Rearrange to obtain the Lineweaver-Burk equation: A plot of 1/v versus 1/[S] should yield a straight line.
Linear Plots Can Be Derived from the Michaelis-Menten Equation The Lineweaver-Burk equation. Figure 13.9 The Lineweaver-Burk double-reciprocal plot.
Enzymatic Activity is Strongly Influenced by pH • Enzyme-substrate recognition and catalysis are greatly dependent on pH. • Enzymes have a variety of ionizable side chains that determine its secondary and tertiary structure and also affect events in the active site. • Substrate may also have ionizable groups. • Enzymes are usually active only over a limited range of pH. • The effects of pH may be due to effects on Km or Vmax or both.
Enzymatic Activity is Strongly Influenced by pH Figure 13.11 The pH activity profiles of four different enzymes.
The Response of Enzymatic Activity to Temperature is Complex • Rates of enzyme-catalyzed reactions generally increase with increasing temperature. • However, at temperatures above 50° to 60° C, enzymes typically show a decline in activity. • Two effects here: • Enzyme rate typically doubles in rate for ever 10°C as long as the enzyme is stable and active. • At higher temperatures, the protein becomes unstable and denaturation occurs.
The Response of Enzymatic Activity to Temperature is Complex Figure 13.12 The effect of temperature on enzyme activity.
13.4 Inhibition of Enzyme Activity • Enzymes may be inhibited reversibly or irreversibly. • Reversible inhibitors may bind at the active site or at some other site. • Reversible inhibitors typically change VM, KM or both. • There are three common types of reversible inhibition: • Competitive • Non-competitive • Uncompetitive