Database Engine Design a.k.a. Research@ DSL
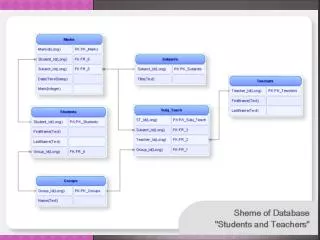
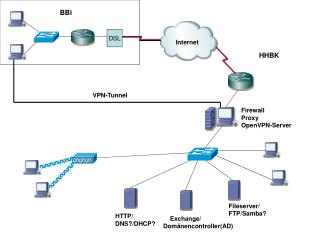
Database Engine Design a.k.a. Research@ DSL . Jayant Haritsa. Database Management Systems (DBMS). Efficient and convenient mechanisms for storing, querying and maintenance of enterprise data Cornerstone of computer industry Uses more than 80 percent of computers worldwide

Database Engine Design a.k.a. Research@ DSL
E N D
Presentation Transcript
Database Engine Designa.k.a. Research@ DSL Jayant Haritsa
Database Management Systems (DBMS) • Efficient and convenient mechanisms for storing, querying and maintenance of enterprise data • Cornerstone of computer industry • Uses more than 80 percent of computers worldwide • Employs more than 70 percent of computer professionals • Largest monetary sector of computer business
DBMS FEATURES • Handle data of arbitrary size • Income-Tax records are in Petabytes (1015) • Self-contained • contains both data and meta-data • Program-Data insulation • application s/w not affected by storage changes SR No | Name | Address | Hostel | GPA SR No | Name | Address | GPA | Hostel
DBMS FEATURES (contd) • Declarative Access • state what you want, not how to get it • On-the-Fly Questions • ask new questions without writing new programs • PEACE OF MIND • changes to the database are guaranteed to be immune to subsequent system failures Sri Sri Ravishankar of the Information World
Current Database Systems • Commercial • IBM DB2 / Oracle / Microsoft SQL Server / Sybase • Public-domain • PostgreSQL / MySQL / Berkeley DB
DBMS Myths • Databases? Isn’t that the boring part of accounting? • Hazaar dumb Cobol programming! • Maha-bore - almost as dull as watching Rahul Dravid bat! • High-tech name for data entry! • Will only get job with TCS! • ...
DBMS Realities • Design of database engines has lots of really, really interesting intellectual problems with practical impact • theory, algorithms, data structures, experiments, prototypes • Turing awards • 1981: Edgar Codd (relational data model) • 1999: Jim Gray (transaction model) • Ullman, Silberschatz, Papadimitrou, … • Rajaraman, Patnaik, Balakrishnan, Jacob/Govindarajan …
Database Systems Lab(DSL) Established 1995
Research Topics • Real-TimeDatabase Systems • Distributed TransactionManagement • OODBMS • Web Databases • Data Mining • XML Databases • Biological Databases • Query Optimization • Multilingual Databases • Music Databases 1995-2000 2000-2005 Last few years
MIDDLEWARE OO Models Mining XML Research Trajectory CORE DB TECHNOLOGY TransactionProcessing AccessMethods Query Processing
Research Techniques • Theory • real-time, data mining, query optimization • Simulation studies • real-time, distributed, web dbms • Empirical evaluation • data mining, biological, multilingual dbms, query optimization • Prototype development • OODBMS (Flexible Manufacturing [MIDAS], VLSI [DIAS], Bio-diversity [Oshadhi,Bodhi] ) • XML (Storage [LegoDB], Compression [XGrind] ) • Query Optimization (Clustering [Plastic], Visualization [Picasso] ) • Multilingual Databases (Cross-lingual SQL [Mira] )
A T CT$ ATTACT$ GTTAATTACT$ A TA TTACT$ CT$ $ 3 4 9 0 7 ATTACT$ 8 CT$ ATTACT$ CT$ 2 6 5 1 Standard Genomic Index: Suffix Tree [Weiner 1973] Vertically-compressed trie of suffixes augmented with links 0 1 2 3 4 5 6 7 8 9 Data =‘GTTAATTACT$’ Suffix Links (xW → W) Tree Edges Search forQuery =‘TTA’ 5 1
Locate all Maximal Matching Substrings[Chang & Lawler 1990] • For each position in query sequence Q, locate all longest matching substrings of length ≥in the indexed data sequence D • Example: D = ‘GTTAATTACT$’Q = ‘CTAATGA’ and = 3 Result: { TAAT:<2,1>AAT:<3,2>}
Maximal Substring Searchwith Suffix Tree Index 0 1 2 3 4 5 6 7 8 9 D =‘GTTAATTACT$’ Q =‘CTAATGA’ =3 A T CT$ ATTACT$ GTTAATTACT$ A TA TTACT$ CT$ $ 3 3 4 9 0 7 ATTACT$ 8 CT$ ATTACT$ CT$ 2 6 2 1 5
Features of Suffix Tree Index • Accurate retrieval • no false negatives (unlike BLAST) • LinearTime Complexity for both Constructionand Search! • because of Suffix-links • Widely used • More than 40-50 applications over biological sequences [Gusfield, 2002] • MUMmer [Celera Genomics], AVID, …
Crippling Limitation • Viable only for sequences that are short enough for their associated suffix tree to fit completely in main memory …[Baeza-Yates and Navarro, 2000] • Best that has been built so far is for sequences of ~ 10 Mbp (Human Genome is 300 times longer!)
Difficulties in Supporting Suffix Trees on Long Sequences - 1 Space overheads are enormous • Order(s) of magnitude larger than data! • Human Genome can be easily stored in main memory (~1 GB) but the index couldbe of the order of 10-100 GB • Disk-resident suffix trees for long sequences
Difficulties in Supporting Suffix Trees on Long Sequences - 2 Tree Construction on Disk is Very Slow • Due to disk thrashing from random seeks The active suffix creeps through the text like a caterpillar … corresponding active node swings through the tree like a butterfly[Giegerich and Kurtz, 1995]
Difficulties in Supporting Suffix Trees on Long Sequences - 3 Searching on Disk is Very Slow • Unbalanced Tree Structure • Shape of tree depends onsequence stochastic properties • “Multi-directional” traversals causes disk thrashing • Tree-Edge “Vertical Walk-Down” • Suffix-Link “Horizontal Jump-Across” Suffix Tree Search • Edge + Link mesh • Two phase Search • Locate • Report ≥ Combination of Batman and Spiderman !
The SPINE* IndexA Horizontally-Compacted Trie Index [*SequenceProcessingINdexing Engine]
0 0 0 A 0 C(0) 1 1 C 2 2 1 A(1) C 1 2 3 3 A 4 4 1(2) C 2 A 5 5 6 C 7 SPINE Index Structure D = ‘ACCACAC’ Complete horizontal compaction into single linear chain!! • Nodes • Forward Edges • Vertebras (Backbone) • Ribs / Ext-Ribs • Backward Edges • Links Root node Link Rib Extension rib Vertebra
0 A 1 C (0) C 2 C A (1) 3 A 4 Structural Advantages of SPINE w.r.t. Suffix Trees • Number of nodes is equal to length of string, whereas in suffix tree can go up to double. • Entire data sequence explicitly embedded in index throw away the data! • On-line incremental algorithm (by definition) • do not need to possess entire data sequence in advance • Node creation order andlogical order are the same prefix-partitionable D =‘ACCA’
0 A 1 C (0) C 2 C A (1) 3 A 4 Advantages of SPINE (contd) • Each node represents a set of suffixes whereas in suffix tree each node represents only a single suffix • Number of suffixes processed for construction and searching is smaller • Easy to develop buffering strategies forpersistent implementations
SPINE Performance Summary Data SetsEcoli: 3.5 Mbp Celegans: 15.5 Mbp HC 21: 28.5 Mbp HC19: 57.5 Mbp Suffix Tree (MUMmer - Celera Genomics) • Spine Space • ~ 2/3 of Suffix Tree • Spine Time • Construction: ~ 1/2 of Suffix Tree • Searching: ~ 1/2 of Suffix Tree
SPINE Summary • First index based on horizontal (inter-path) compaction of the trie • Collapses into a single linear structure • Improved features and performance w.r.t. suffix trees, the classical index • Prefix-partitionable (first index to have this property) • Easily amenable to persistent disk implementation • Retains linear time/space complexity • Better construction speed and capacity • Better search response times