V5: mRNA translation
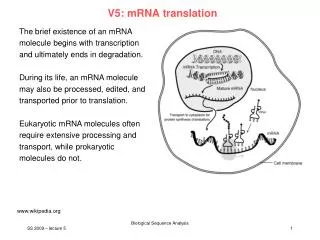
V5: mRNA translation. The brief existence of an mRNA molecule begins with transcription and ultimately ends in degradation. During its life, an mRNA molecule may also be processed, edited, and transported prior to translation.

V5: mRNA translation
E N D
Presentation Transcript
V5: mRNA translation The brief existence of an mRNA molecule begins with transcription and ultimately ends in degradation. During its life, an mRNA molecule may also be processed, edited, and transported prior to translation. Eukaryotic mRNA molecules often require extensive processing and transport, while prokaryotic molecules do not. www.wikipedia.org SS 2009 – lecture 5 Biological Sequence Analysis 1
eukaryotic pre-mRNA processing The short-lived, unprocessed or partially processed, product of transcription by RNA polymerase is termed pre-mRNA. Once completely processed, it is termed mature mRNA. Processing of mRNA differs greatly among eukaryotes, bacteria and archea. Non-eukaryotic mRNA is essentially mature upon transcription and requires no processing, except in rare cases. Eukaryotic pre-mRNA, however, requires extensive processing. www.wikipedia.org SS 2009 – lecture 5 Biological Sequence Analysis 2
processing: 5‘ cap addition A 5' cap is a modified guanine nucleotide that has been added to the 5' end of a eukaryotic messenger RNA shortly after the start of transcription. The 5' cap consists of a terminal 7-methylguanosine residue which is linked through a 5'-5'-triphosphate bond to the first transcribed nucleotide. Its presence is critical for recognition by the ribosome, pre-mRNA splicing, 3‘end formation, U snRNA transport, and regulation of decay. Cap addition is coupled to transcription, and occurs co-transcriptionally, such that each influences the other. Shortly after the start of transcription, the 5' end of the mRNA being synthesized is bound by a cap-synthesizing complex associated with RNA polymerase. www.wikipedia.org m7GpppG binding to nuclear cap-binding protein complex SS 2009 – lecture 5 Biological Sequence Analysis 3 Calero et al. Nat. Struct. Biol. 9, 912 (2002)
processing: editing In some instances, an mRNA will be edited, changing the nucleotide composition of that mRNA. mRNA has been observed in tRNA, rRNA, and mRNA molecules of eukaryotes but not prokaryotes. RNA editing mechanisms include nucleoside modifications such as C to U and A to I deaminations, as well as non-templated nucleotide additions and insertions. RNA editing alters the amino acid sequence of the encoded protein so that its sequence differs from that predicted from the genomic DNA sequence. An example in humans is the apolipoprotein B mRNA, which is edited in some tissues, but not others. Here, the editing creates an early stop codon, which upon translation, produces a shorter protein. www.wikipedia.org SS 2009 – lecture 5 Biological Sequence Analysis 4
processing: polyadenylation • Polyadenylation is the covalent linkage of a polyadenylyl moiety to an mRNA molecule. In eukaryotic organisms, most mRNA molecules are polyadenylated at the 3' end. • The poly(A) tail and the protein bound to it aid in protecting mRNA fromdegradation by exonucleases. • Polyadenylation is also important for • transcription termination, • export of the mRNA from the nucleus, and • translation. www.wikipedia.org SS 2009 – lecture 5 Biological Sequence Analysis 5
processing: polyadenylation Polyadenylation occurs during and immediately after transcription of DNA into RNA. After transcription has been terminated, the mRNA chain is cleaved through the action of an endonuclease complex associated with RNA polymerase. After the mRNA has been cleaved, around 250 adenosine residues are added to the free 3' end at the cleavage site. This reaction is catalyzed by polyadenylate polymerase. Just as in alternative splicing, there can be more than one polyadenylation variant of a mRNA. www.wikipedia.org SS 2009 – lecture 5 Biological Sequence Analysis 6
RNA splicing Splicing is the process by which pre-mRNA is modified to remove certain stretches of non-coding sequences called introns; the stretches that remain include protein-coding sequences and are called exons. This is needed for the typical eukaryotic messenger RNA before it can be used to produce a correct protein through translation. For many eukaryotic introns, splicing is done in a series of reactions which are catalyzed by the spliceosome, a complex of small nuclear ribonucleoproteins (snRNPs), but some RNA molecules are also capable of catalyzing their own splicing (ribozymes). www.wikipedia.org
splicing repression There are 2 major types of RNA sequence elements present in pre-mRNAs, and specific RNA-binding proteins bind to each of these elements. Silencers are sites to which splicing repressor proteins bind, reducing the probability that a nearby site will be used as a splice junction. These can be located in the intron itself (intronic splicing silencer, ISS) or in a neighboring exon (exonic splicing silencer, ESS). They vary in sequence, as well as in the types of proteins that bind to them. The majority of the repressors that bind are heterogeneous nuclear ribonucleo-proteins (hnRNPs) such as hnRNPA1 and polypyrimidine tract binding protein (PTB). www.wikipedia.org SS 2009 – lecture 5 Biological Sequence Analysis 8
splicing activation Splicing Enhancers are sites to which splicing activator proteins bind, increasing the probability that a nearby site will be used as a splice junction. Splicing enhancers also may occur in the intron (intronic splicing enhancer, ISE) or exon (exonic splicing enhancer, ESE). Most of the activator proteins that bind to ISEs and ESEs are members of the SR protein family. Such proteins contain RNA recognition motifs and arginine and serine-rich (RS) domains. www.wikipedia.org SS 2009 – lecture 5 Biological Sequence Analysis 9
alternative splicing (AS) Alternative splicing is a RNA splicing variation mechanism in which the exons of the primary gene transcript, the pre-mRNA, are separated and reconnected so as to produce alternative ribonucleotide arrangements. These linear combinations are then translated into different proteins. In this way, AS uses genetic expression to facilitate the synthesis of a greater variety of proteins. www.wikipedia.org SS 2009 – lecture 5 Biological Sequence Analysis 10
5 basic modes of alternative splicing (1) Exon skipping: here, an exon may be spliced out of the primary transcript or retained. This is generally the most common mode in mammalian pre-mRNAs. (2) Mutually exclusive exons: One of two exons is retained in mRNAs after splicing, but not both. (3) Alternative donor site: An alternative 5' splice junction (donor site) is used, changing the 3' boundary of the upstream exon. (4) Alternative acceptor site: An alternative 3' splice junction (acceptor site) is used, changing the 5' boundary of the downstream exon. (5) Intron retention: A sequence may be spliced out as an intron or simply retained. This is distinguished from exon skipping because the retained sequence is not flanked by introns. If the retained intron is in the coding region, the intron must encode amino acids in frame with the neighboring exons, or a stop codon or a shift in the reading frame will cause the protein to be non-functional. This is generally the rarest mode in mammals. www.wikipedia.org SS 2009 – lecture 5 Biological Sequence Analysis 11
Example: alternative splicing of Drosophila dsx pre-mRNA Pre-mRNAs from the D. melanogaster gene dsx contain 6 exons. In males, exons 1,2,3,5,and 6 are joined to form the mRNA, which encodes a trans-criptional regulatory protein required for male development. In females, exons 1,2,3, and 4 are joined, and a poly-A signal in exon 4 causes cleavage of the mRNA at that point. The resulting mRNA is a transcriptional regulatory protein required for female development. De-Leon, Annu Rev. Biophys Biomol Struct. 36, 191 (2007) SS 2009 – lecture 5 Biological Sequence Analysis 12
importance of alternative splicing Alternative splicing is of great importance to genetics - it invalidates the old "one-gene-one-protein" hypothesis. External information is needed in order to decide which polypeptide is produced, given a DNA sequence and pre-mRNA. Possibly, this was a very important step towards higher efficiency of eukaryotic genomes, because information can be stored much more economically. Several proteins can be encoded in a DNA sequence whose length would only be enough for two proteins in the prokaryote way of coding. Alternatively, a new protein can evolve without changing the DNA of a gene. Instead, the same effect can be achieved by differential regulation. De-Leon, Annu Rev. Biophys Biomol Struct. 36, 191 (2007) SS 2009 – lecture 5 Biological Sequence Analysis 13
importance of alternative splicing Humans have only about twice as many genes as Caenorhabditis elegans or the fly Drosophila melanogaster. How can one explain the greater complexity of humans? Hypothesis: The greater complexity of humans, or vertebrates generally, might be due to higher rates of AS in humans than in invertebrates. However, EST studies showed that the frequency of AS is similar in human to that in mouse, rat, cow, fly, worm, and the plant Arabidopsis thaliana. The "record-holder" for AS is a D. melanogaster gene called Dscam, which has 38,016 splice variants. De-Leon, Annu Rev. Biophys Biomol Struct. 36, 191 (2007) SS 2009 – lecture 5 Biological Sequence Analysis 14
Example of AS: Transient Receptor Potential channels • The general topology of a TRP subunit consists of • 6 predicted TM domains with • a putative pore loop between TMD5 and TMD6 and • intracellular N- and C-terminal regions of variable length, • the former containing multiple ankyrin (ANK) repeats in the TRPC, TRPA, TRPN, and TRPV subfamilies. • Functional TRP channels are supposed to result following the assembly of 4 TRP subunits. Arniges et al. J. Biol. Chem. 281, 1580 (2006)
Identification of TRMP3 splice variants from mouse brain A, schematic diagram of the mouse Trpm3 gene, comprising 28 exons. Oberwinkler et al. J. Biol. Chem. 280, 22540 (2005)
TRPM3 splice variants C, schematic presentation of TRPM3 with transmembrane domains 1–6, coiled coil region (cc), and TRP homology domain (Trp). Novel mouse TRPM3 protein variants shown as thick black lines are compared with the human variants hTRPM3a–f and hTRPM31325. The numbers of amino acid residues of each variant are indicated in parentheses. Starting from residue 156, mouse and human TRPM3 have 97% sequence identity. Oberwinkler et al. J. Biol. Chem. 280, 22540 (2005)
Pore regions of splice variants D, putative pore regions of TRPM31 and TRPM32 compared with the corresponding mouse sequences of TRPM6, TRPM7, TRPV5, and TRPV6. The 12 additional amino acid residues present in TRPM31 are indicated. Identical residues are boxed in black, conserved in gray. An Asp residue that determines Ca2+ permeation of the TRPV5/TRPV6 pore is marked by an asterisk. Residues proposed to build the selectivity filter of TRPV6 are underlined. Oberwinkler et al. J. Biol. Chem. 280, 22540 (2005)
TRPM3 functions as cation channel Heterologous expression of TRPM31 induces outwardly rectifying cation currents inhibited by intracellular Mg2+. A,current-voltage relationship of a TRPM31-expressing cell in standard Ringer or NMDG solution within 60 s after establishing the whole cell patch clamp configuration. Oberwinkler et al. J. Biol. Chem. 280, 22540 (2005)
Permeability for divalent cations TRPM31 and TRPM32 display large differences in their relative permeability ratios for divalent cations. A, comparison of TRPM31 and TRPM32 currents at 80 mV and 80 mV in extracellular solutions containing indicated amounts of Ca2+. B, reversal potential during the experiment shown in panel A. Oberwinkler et al. J. Biol. Chem. 280, 22540 (2005)
Identification of TRMP3 variants from mouse brain C and D, statistical analysis of reversal potential measurements in experiments similar to that shown in panel B during the application of solutions containing the indicated concentration of Ca2+ (C) or Mg2+ (D) as the only permeable ion. Continuous thin lines show the expected reversal potential calculated from Goldman-Hodgkin-Katz theory for the indicated relative permeability ratios. Each point represents the mean of 3–15 independent measurements (at a divalent concentration of 10 mM p < 0.001, otherwise at least p < 0.05). Oberwinkler et al. J. Biol. Chem. 280, 22540 (2005)
Effect of extracellular cations Inhibition of TRPM3-dependent currents by extracellular cations. A, comparison of TRPM31 and TRPM32 currents at 80 mV and 80 mV in extra-cellular solutions containing indicated amounts of Na+. Outward currents through TRPM31 are unaffected by extracellular Na+, whereas outward currents through TRPM32 are inhibited in a dose-dependent manner by these ions. B, statistical analysis of recordings with varying concentrations of Na+, K+, Ca2+, and Mg2+. • TRPM32 is inhibited by all cations tested on the extra- cellular side. Oberwinkler et al. J. Biol. Chem. 280, 22540 (2005)
Summary Alternative Splicing Switches the Ion Selectivity of TRPM3 Channels— The selectivity of ion channels is thought to be determined by the geometry and charge distribution of the selectivity filter, usually envisioned as the narrowest part of the channel pore. Typically, all members of an ion channel family, such as voltage-gated Na+, K+, or Ca2+ channels, share common ionic selectivities. Oberwinkler et al. J. Biol. Chem. 280, 22540 (2005)
Summary II The TRP family of ion channels is unusual in this respect as its members have quite diverging cationic selectivity profiles. The Trpm3 gene adds extra complexity to this picture, because two channels can be expressed from this gene with entirely different ionic selectivities. One channel, TRPM31, preferentially conducts monovalent cation influx, whereas TRPM32 strongly favors divalent entry. In vivo, such a change in ionic selectivity must be expected to have considerable consequences for the function of the channel and the physiology of the cell that expresses it. The switch of ionic selectivity in TRPM3 variants is due to removal of a short stretch of 12 amino acid residues and exchanging 1 further residue within the linker domain between the presumed fifth and sixth transmembrane regions. Oberwinkler et al. J. Biol. Chem. 280, 22540 (2005)
Summary III Compared with the presumed pore regions of other members of the TRP family, the pore loop of TRPM3 is considerably longer by 8 (TRPM32) and 20 (TRPM31) additional amino acid residues. The domains that build the proposed selectivity filter of the Ca2+-selective TRPV5/V6 channels are conserved in TRPM3 proteins. The splicing within the TRPM3 channel pore introduces additional, positively charged amino acid residues into this domain. This might decrease the Ca2+ permeability of TRPM31 compared with TRPM32, perhaps simply because of increased electrostatic repulsion. Block of TRPM3 Channels by Intra- and Extracellular Cations — Both TRPM31 and TRPM32 are regulated by physiological concentrations of intracellular Mg2+, similar to related members of the TRPM family such as TRPM6 and TRPM7. Oberwinkler et al. J. Biol. Chem. 280, 22540 (2005)
end of translation: action of ribosome www.wikipedia.org SS 2009 – lecture 5 Biological Sequence Analysis 26
additional slides SS 2009 – lecture 5 Biological Sequence Analysis 27
TRPV4 channels The non-selective cation channel TRPV4 is a member of the transient receptor potential (TRP) family of channels. TRPV4 shows multiple modes of activation and regulatory sites, enabling it to respond to various stimuli, including osmotic cell swelling, mechanical stress, heat, acidic pH, endogenous ligands, high viscous solutions, and synthetic agonists such as 4-phorbol 12,13-didecanoate. TRPV4 mRNA is expressed in a broad range of tissues, although functional tests have only been carried out in a few: endothelial, epithelial, smooth muscle, keratinocytes, and DRG neurons. Arniges et al. J. Biol. Chem. 281, 1580 (2006)
Cloning of TRPV4 variants from human airway epithelial cells • A reverse transcriptase-PCR-based cloning process identified 5 variants of the TRPV4 channel in human tracheal epithelial cells. • 2 of the cloned cDNAs corresponded to the already described • TRPV4 isoform A (fulllength cDNA) and • TRPV4 isoform B (lacking exon number 7, 384–444 amino acids). • We also identified 3 new splice variants affecting the cytoplasmic N-terminal region. • TRPV4-C lacks exon 5 (237–284 amino acids), • TRPV4-D presents a short deletion inside exon 2 (27–61 amino acids), and TRPV4-E (237–284 and 384–444 amino acids) is produced by a double alternative splicing lacking exons 5 and 7. Arniges et al. J. Biol. Chem. 281, 1580 (2006)
Different splice variants of TRPV4 A, schematic diagram showing the intracellular N-terminal region of the human TRPV4 channel (amino acids 1–471). Exons and the corresponding amino acids lost in each TRPV4 isoform are indicated by numbers. Arniges et al. J. Biol. Chem. 281, 1580 (2006)
Functional analysis of TRPV4 variants: intracellular [Ca2+] The TRPV4-A channel responds to a wide variety of stimuli. Here, HeLa cells were transiently transfected and intracellular Calcium concentration was determined via Fura-2 ratios as reponse to 3 well known activators of TRPV4-A: 30% hypotonic solution, 1 M 4-PDD, or 10 M arachidonic acid Only TRPV4-A and TRPV4-D show channel activity. Arniges et al. J. Biol. Chem. 281, 1580 (2006)
TRPV4-A and D produce functional channels TRPV4-A and TRPV4-D isoforms produce functional channels with similar properties when expressed in HEK-293 cells. A, current traces obtained from TRPV4-A and TRPV4-D-expressing HEK-293 cells at the indicated voltages in the presence of 1M 4-PDD. Dashed lines indicate the zero current level. B, I–V relationship of 4-PDD-activated TRPV4-A (open circle) and TRPV4-D (closed circle) channels in inside-out patches. Arniges et al. J. Biol. Chem. 281, 1580 (2006)
Retention in ER Co-localization experiments (not shown): TRPV4-B, C and E are trapped in the ER and not translocated to the plasma membrane. Arniges et al. J. Biol. Chem. 281, 1580 (2006)
Homomerization of TRPV4 variants FRET efficiencies determined between identical CFP- and YFP-fused TRPV4 variants (A–E) transiently cotransfected in HEK-293 cells. High FRET efficiencies corresponding to homomultimer formation could only be demonstrated for TRPV4-A and TRPV4-D variants. Arniges et al. J. Biol. Chem. 281, 1580 (2006)
Heteromerization of TRPV4 variants B, FRET efficiencies determined between different TRPV4 variants showed heterooligomerization only for A and D proteins. Arniges et al. J. Biol. Chem. 281, 1580 (2006)
Summary This study of oligomerization, localization, and channel activity of human TRPV4 splice variant identified the N-terminal ANK repeats as key molecular determinants of subunit assembly and subsequent processing of the assembled channel. Five TRPV4 variants (TRPV4-A–E) cloned from human airway epithelial cells were grouped into two classes: group I: TRPV4-A and TRPV4-D group II: TRPV4-B, TRPV4-C, and TRPV4-E. Group I variants are correctly processed and targeted to the plasma membrane where they form functional channels with similar electrophysiological properties. Variants from group II, which are lacking parts of the ANK domains are unable to oligomerize and were retained intracellularly, in the ER. Arniges et al. J. Biol. Chem. 281, 1580 (2006)
Summary II • Discovery of three important traits of TRPV4 biogenesis. • Glycosylation of TRPV4 channel involves ER to Golgi transport with the corresponding change in the N-linked oligosaccharides from the high mannose type characteristic of the ER to the complex type characteristic of the Golgi apparatus, without apparent O-glycosylation. • 2) TRPV4-A subunits oligomerize in the ER. • 3) Impaired subunit assembly of type II variants is because of the lack of N-terminal ANK domains and causes protein retention in the ER. Arniges et al. J. Biol. Chem. 281, 1580 (2006)
Summary III Ion channel functional diversity is greatly enlarged by both the presence of splice variants and heteromerization of different pore-forming and regulatory subunits. Alternative splicing is a major contributor to protein diversity. Within the TRP family of ion channels several splice variants have been identified, some of them resulting in lack of responses to typical stimuli, others modifying the pore properties, and those exerting dominant negative effects. Group II TRPV4 splice variants have been identified in two unrelated, human airway epithelial cell lines. Considering the relevance of TRPV4 channels in epithelial physiology, a change in the expressed ratio of group I to group II variants, favoring the later, may modify normal epithelial functioning. Splicing can be regulated by several stressing stimuli including pH, osmotic, and temperature shocks, all of them being also activating stimuli of the TRPV4. Arniges et al. J. Biol. Chem. 281, 1580 (2006)
mRNA translation In activation, the correct amino acid is covalently bonded to the correct transfer RNA (tRNA). While this is not technically a step in translation, it is required for translation to proceed. The amino acid is joined by its carboxyl group to the 3' OH of the tRNA by an ester bond. When the tRNA has an amino acid linked to it, it is termed "charged". Initiation involves the small subunit of the ribosome binding to 5' end of mRNA with the help of initiation factors (IF). Termination of the polypeptide happens when the A site of the ribosome faces a stop codon (UAA, UAG, or UGA). When this happens, no tRNA can recognize it, but a releasing factor can recognize nonsense codons and causes the release of the polypeptide chain. The 5' end of the mRNA gives rise to the proteins N-terminal and the direction of translation can therefore be stated as N->C. De-Leon, Annu Rev. Biophys Biomol Struct. 36, 191 (2007) SS 2009 – lecture 5 Biological Sequence Analysis 39
mRNA translation The mRNA carries genetic information encoded as a ribonucleotide sequence from the chromosomes to the ribosomes. The ribonucleotides are "read" by translational machinery in a sequence of nucleotide triplets called codons. Each of those triplets codes for a specific amino acid. The ribosome and tRNA molecules translate this code to a specific sequence of amino acids. The ribosome is a multisubunit structure containing rRNA and proteins. It is the "factory" where amino acids are assembled into proteins. tRNAs are small noncoding RNA chains (74-93 nucleotides) that transport amino acids to the ribosome. tRNAs have a site for amino acid attachment, and a site called an anticodon. The anticodon is an RNA triplet complementary to the mRNA triplet that codes for their cargo amino acid. De-Leon, Annu Rev. Biophys Biomol Struct. 36, 191 (2007) SS 2009 – lecture 5 Biological Sequence Analysis 40
mRNA translation Aminoacyl tRNA synthetase catalyzes the bonding between specific tRNAs and the amino acids that their anticodons sequences call for. The product of this reaction is an aminoacyl-tRNA molecule. This aminoacyl-tRNA travels inside the ribosome, where mRNA codons are matched through complementary base pairing to specific tRNA anticodons. The amino acids that the tRNAs carry are then used to assemble a protein. The energy required for translation of proteins is significant. For a protein containing n amino acids, the number of high-energy Phosphate bonds required to translate it is 4n-1. The rate of translation varies; it is significantly higher in prokaryotic cells (up to 17-21 amino acid residues per second) than in eukaryotic cells (up to 6-7 amino acid residues per second) De-Leon, Annu Rev. Biophys Biomol Struct. 36, 191 (2007) SS 2009 – lecture 5 Biological Sequence Analysis 41