Molecular Genetics - Structure
DESCRIPTION
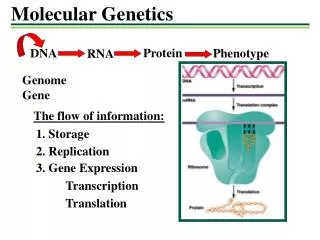
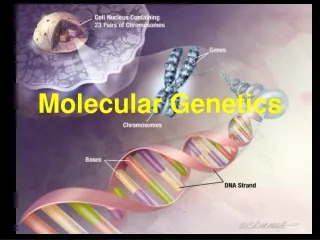
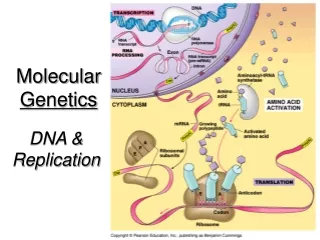
Molecular Genetics - Structure. Chapter 4. History – 4.1. DNA – deoxyribonucleic acid is a double stranded polymer of nucleotides (sugar, phosphate and one of four nitrogenous bases) that carry genetic information of an organism. How did we get to an understanding of this?.
1 / 0
Télécharger la présentation 

Molecular Genetics - Structure
An Image/Link below is provided (as is) to download presentation
Download Policy: Content on the Website is provided to you AS IS for your information and personal use and may not be sold / licensed / shared on other websites without getting consent from its author.
Content is provided to you AS IS for your information and personal use only.
Download presentation by click this link.
While downloading, if for some reason you are not able to download a presentation, the publisher may have deleted the file from their server.
During download, if you can't get a presentation, the file might be deleted by the publisher.
E N D
Presentation Transcript
-
Molecular Genetics - Structure
Chapter 4 - History – 4.1 DNA – deoxyribonucleic acid is a double stranded polymer of nucleotides (sugar, phosphate and one of four nitrogenous bases) that carry genetic information of an organism. How did we get to an understanding of this?
- History/Experiments Mendel's work with peas and Morgan's work with Drosophilia demonstrated that traits can be transmitted from parent to offspring in different combinations and an overall realization of the science of genetics began around the turn of the twentieth century. Late 1860's – Miescher isolated non-protein substance from the nucleus of cells; named this substance nuclein. At this point proteins were believed to be the main hereditary material
- History/Experiments 1920's - Levene worked out the basic chemistry of nucleic acids which consisted of: Phosphate (PO4) groups 5-C sugars (deoxyribose, no –OH on the 2’; ribose, -OH) 4 nitrogenous bases: A, G = double; C,T,U = single Three factors in roughly equal proportions so DNA/RNA molecules must have all three strung together in chains. Each unit is called a nucleotide with the nitrogenous base being the only factor that differentiates
- History/Experiments
- History/Experiments Putting them together One end of the DNA chain always ends with a phosphate group that is attached to the number 5 carbon of the deoxyribose sugar (5' end) The other end of the DNA chain terminates with a hydroxyl group that is attached to the number 3 carbon of the deoxyribose sugar (3' end) The chemical reaction between the phosphate group of one unit and the hydroxyl group of the other causes the elimination of water to form a covalent bond (phosphodiester bond). That is two ester bonds (-O-) with phosphorous in-between. The nitrogenous base is attached to the 1’ carbon of the sugar by a glycosyl bond (bond between a sugar and another organic molecule by way of an intervening N or O atom).
- History/Experiments
- History/Experiments 1920's - Griffith's experiments with Streptococcus pneumoniae Heat killed pathogenic inserted into mice = live mice Live pathogenic strain inserted into mice = mice die (due to pathogenic capsule) Live non-pathogenic strain inserted into mice = mice live However Mixture of heat killed pathogenic and live non-pathogenic = mice die???? both should have been harmless but the live cells have been "transformed" by the dead ones, that is, genetic information specifying the capsule of the heat killed had passed from the dead cells to the living ones Avery added to this experiment by analyzing the "transforming principle" and noticed that it was not destroyed by protein destroying enzymes but was destroyed by DNAase
- History/Experiments 1930s - Hammerlings work lead to a likely understanding of where the genetic material was stored in the cell Acetabulariacrenulata (one celled green algae) and A. mediterranea and using grafting techniques. The nucleus is what determined the final phenotype of the aquatic plant. 1941 - Beadle and Tatum - One Gene/One Enzyme Hypothesis
- History/Experiments 1949 - Chargaff showed that the 4 main bases for DNA were not present in equal amounts and that it varied in significant ways. A=T and G=C known as Chargaff's rule.
- History/Experiments 1952 - Hershey-Chase
- History/Experiments 1953 – Rosalind Franklin was working with X-ray crystallography to determine the structure of DNA Biochemist Maurice Wilkins understood how to get a more definite understanding of what the diffraction of the X-ray meant and took the findings, that the structure had to have a corkscrew orientation about 2nm in diameter and one complete helical turn every 3.4nm, to Watson and Crick (see below). She deduced that DNA had to be helical with a steady state of turn through the structure and had photographs to prove it. Two young guys by the names of Watson and Crick caught wind of this, found out the information that had been collected and quickly put together a model through deductive reasoning that is the double helix model that we know of today. They noted that DNA consists of two antiparallel strands of nucleotides. That means that the two strands are parallel but run in opposite directions (the 5’ end of one strand of DNA aligns with the 3’ end of the other strand). Also, the bases pointed inward and were bonded together, purine from one helix with a pyrimidine from the other, with H bonds, which nicely supported Chargaff's rule to add to their argument. This is referred to as complementary base pairing.
- History/Experiments 1958 - Meselson and Stahl looked at the complementarities of DNA and its replication, that is, any new DNA that is created is an exact compliment of the old strand They grew bacteria in heavy 15N for 17 generations and then transferred to 14N and harvested the bacteria at various intervals. F1 was an intermediate band and F2 had the intermediate and a 14N band (DNA that included none of the heavy band). Supports the idea that the individual stands remain intact during DNA replication when they serve as templates for new DNA strands, and after replication, each strand is found paired with a new complementary strand Applied to bacteria at this point however eukaryotic cells follow the same pattern of semi-conservative replication.
- History/Experiments
- DNA Replication and Repair - 4.3 DNA Replication - General Proteins bind to a specific site on the DNA known as the replication origin (eukaryotes have many) Enzyme (DNA helicase) unwinds the double helix by breaking the H bonds. Single stranded binding proteins (SSBs) bind to the exposed DNA single strands and do not allow them to anneal. Enzyme DNA gyrase relieves tension on bacterial DNA…similar enzyme present in eukaryotic cells. Replication occurs in two directions from the point of origin (replication fork)
- DNA Replication
- DNA Replication DNA Replication - Specific Enzyme DNA polymerase III synthesizes complementary strands of DNA during DNA replication Works 5’ to 3’ so it adds deoxyribonucleoside triphosphates (deoxyribose bonded to three phosphate groups and a nitrogenous base) to a 3’ end of the leading strand.
- DNA Replication Enzyme primase creates RNA primer which anneals to a 10 to 60 bp section of the template strand. Once this RNA primer is in place DNA polymerase III can start the elongation process. The energy for the process is partially driven by breaking off the first and second phosphate groups and this allows the complementary nucleotide to be attached via dehydration synthesis
- DNA Replication Leading strand uses 3’ to 5’ template and is built toward the replication fork Lagging strand uses the 5’ to 3’ template and therefore many short fragments are created called Okazaki fragments. DNA polymerase I removes the RNA primers and replaces them with appropriate deoxyribonucleotides. DNA ligase uses phosphodiester bonds to link Okazaki fragments
- DNA Replication
- DNA Repair DNA polymerase I & III act as quality control and proofread new sequences. When a mistake is found either enzyme acts as an exonuclease. It cuts out the inappropriate nucleotide and adds in the appropriate one.
More Related

Audio
Live Player