Systemic acquired resistance
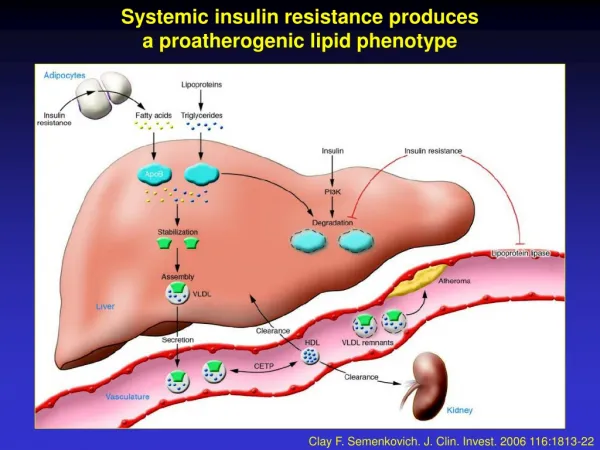
Systemic acquired resistance. Systemic acquired resistance was first recognized as a significant phenomenon in 1933 by Chester. He and many others since note that infection of plants with necrotizing pathogens (causing HR)

Systemic acquired resistance
E N D
Presentation Transcript
Systemic acquired resistance Systemic acquired resistance was first recognized as a significant phenomenon in 1933 by Chester. He and many others since note that infection of plants with necrotizing pathogens (causing HR) often results in enhanced resistance to subsequent infections by a variety of fungal, bacterial and viral pathogens. This physiological immunity was termed systemic acquired resistance (SAR). SAR confers a broad spectrum type of resistance SAR is effective against some but not all pathogens: Tobacco: Phytophthora parasitica, Cercospora nicotianae, Peronospora tabacina tobacco mosaic virus, tobacco necrosis virus, Pseudomonas syringae pv. tabaci, Erwinia carotovora Not effective against: Botrytis cinerea or Alternaria alternata Arabidopsis: Phytophthora parasitica turnip crinkle virus Pseudomonas syringae pv. tomato DC3000
Some key events in understanding regulation of SAR Systemic acquired resistance was associated with the coordinated induction of a set of SAR genes encoding proteins known as Pathogenesis-related (PR) proteins (Van Loon and Gianinazzi (early 1970s). (1979) White found that acetyl salicylic acid application sufficient to induce PR gene expression and enhanced resistance to tobacco mosaic virus in tobacco plant. Discovery came out the interest in developing chemical control methods for viral infection. After that several groups went on to show that salicylic acid application on tobacco leaves mimics pathogen induced expression of PR genes and pathogen resistance in treated tissues. (1990) Two groups one led by Klessig and Raskin and another led by Metraux found that salicylic acid accumulates in cucumber and tobacco plants prior to pathogen infection, but before the onset of resistance. The work by these and many others led to the hypothesis that salicylic acid (SA) is the endogenous signal molecule that is required for the induction of systemic acquired resistance. (1993/1994) The group headed by Ryals made tobacco plants that could not accumulate SA and found that these plants were defective in their ability to develop systemic acquired resistance. This work demonstrated a central role for SA in establishing systemic acquired resistance. The group also demonstrated that these tobacco plants were defective in their ability to accumulate PR proteins. (1997) Cloning of NPR1, a key regulator of SAR
PR (Pathogenesis-Related) proteins PR proteins first identified as major proteins induced by necrotizing pathogens (pathogens that induced the hypersensitive response) • Proteins secreted predominantly into intercellular spaces in response to wounding or infection. • Soluble at pH 3 • Basic homologs also found (in vacuole). • Proteinase resistant (but not proteinase inhibitors). • Some are developmentally expressed as part of normal plant development in absence of wound or infection (e.g. flowering).
Traditional PR protein gels Acidic gel Basic gel PR proteins Proteins first isolatedfrom apoplast ofTMV-infected tobacco. Induced by many other pathogens. Some PR proteins are also induced byabiotic stresses. old nomenclature All tobaccoPR proteins
What do PR proteins do? Type Family member Biochemical function Resistance PR-1a,b,c Tob 1a Unknown Oomycetes (1a) PR-2 Tob. N, O 1,3 -glucanase antifungal (w/ chitinase) PR-3 Tob. P, Q chitinases anti-Rhizoctonia, other fungi PR-4 Tob. R Unknown antifungal PR-5 Tob. S Homologous to amylase /proteinase inhibitor and sweet protein thaumatin. Anti-oomycete PR-6 Tom. Inhib I proteinase inhibitor PR-7 Tom. P69 endoproteinase PR-8 Cuc. Chitinase chitinase PR-9 Tob. Lignin- peroxidase forming peroxidase PR-10 Parsely PR-1 ribonuclease-like PR-11 Tob. V chitinase chitinase PR-12 Defensins antifungal PR-13 Thionins antifungal PR-14 Lipid-transfer proteins antifungal Modified from Table 21.4, Biochem & Mol Biol of Plants
Chitin is 1,4 N-acetyl-D-glucosamine (GlcNAc) polymer Chitosan is partially deacetylated chitin.
Control Line 373 230 238 329 373 548 18 days after growth in R. solani-laden sand 11 d.a.g. in R. solani sand Constitutive expression of chitinase PR protein confersresistance to Rhizoctonia solani Broglie et al. (1991) Science 254, 1194-1196
Application of salicylic acid mimics SAR (1979) White found that the application of aspirin, salicylic acid, and benzoic acid resulted in enhanced resistance to TMV. Used 3 tobacco cultivars that contained the N resistance gene that confers HR to TMV. Found > 90% reduction in lesion number in treated leaves versus water control.
SA accumulation is associated with acquisition of resistance SA Lesions obtained after second inoculation
Central role of SA in SAR PR-1 PR-2 PR-3
Enhanced susceptibility Loss of resistance INA induces resistance in presence of nahG
The Arabidopsis NPR1 Gene That Controls Systemic Acquired Resistance Encodes a Novel Protein Containing Ankyrin Repeats Hui Cao, Jane Glazebrook,Joseph D. Clarke, Sigrid Volko,and Xinnian Dong (1997) Cell 88, 57–63, Signaling steps between SA and PR protein expressionand disease resistance. • Previously: Linked a PR protein promoter called BGL2 to GUS. • Screened thousands of mutant transgenic BGL2-GUS plants forABSENCE of GUS activity induced by SA treatment. • Using standard Arabidopsis genetic mapping methods, identified asingle mutant gene, npr1. Phenotype: • Complete absence of GUS activity in response to SA • Absence of PR-1, PR-5, BGL2 expression in response to SA • Is now susceptible to Peronospora parasitica • and to Pseudomonas syringae pv maculicola (Psm). Cao et al. (1997) Cell 88, 57–63,
genotype: wt npr1-2 Cloned NPR1 by standard 1990’s methods. Chromosome walking, YAC library… non-compl. transgene: none none NPR1 NPR1 Proof of cloning by transgenic comple-mentation of mutants w/ wildtype NPR1. wt npr1-1 + NPR1 npr1-1 symptoms Psm inoculated GUS Cao et al. (1997) Cell 88, 57–63,
NPR1 has ankyrin repeats Ankyrin repeats arein lots of differentproteins. Involved in protein-protein interactions. Especially in proteinsthat control trans-cription.In NF-kB and I-kB inmammals. Inducedby many pathogens, stresses… Cao et al. (1997) Cell 88, 57–63,
NPR1 is reduced to a monomer during plant defense Mou et al., (2003) Cell, 113:935–944
Expression of PR-1 is associated with NPR1 monomerization Mou et al., (2003) Cell, 113:935–944
Monomeric NPR1 localizes to the nucleus Mou et al., (2003) Cell, 113:935–944
NPR1 enhances TGA1 binding to the as-1 element under reducing conditions Despres et al. (2003) Plant Cell. 15:2181–2191,
Identification and Cloning of a Negative Regulator of Systemic Acquired Resistance, SNI1, through a Screen for Suppressors of npr1-1 Xin Li, Yuelin Zhang, Joseph D. Clarke, Yan Li,† and Xinnian Dong* Cell, 98, 329–339, 1999. Screened for EMS mutants of npr1-1 plants containing BGL2-GUSreporter. Look for plants that turn blue in response to INA (SA analog) like NPR1 wild type plants. But which, of course still harbor the npr1-1 mutation. Found 11 loci that gave increased GUS, out of 7000 plants screened. Li et al., Cell 98, 329–339.
SNI1 is similar to mouse Retinoblastoma (Rb). Rb is a tumor suppressor that represses function of E2F transcription factor Li et al., Cell 98, 329–339.
Model for control of PR gene expression by NPR1 and SNI1. SNI1 is in the nucleus nucleus TGA TFs Li et al., Cell 98, 329–339.
Everyone interested in regulation of gene expression in plant defense should know about WRKY transcription factors • Recognize the motif: (T)(T)TGAC(C/T). • Have the conserved WRKYGQK at N-terminal end. • Have a novel zinc-finger-like motif. • Bind DNA via divalent cation (probably zinc). • Approx. 100 members of WRKY family in Arabidopsis. • NPR1 has a WRKY motif in its promoter: TTGACTTGACTTGGCTCTGCTCGTCAA The WRKY superfamily of plant transcription factors Thomas Eulgem, Paul J. Rushton, Silke Robatzek and Imre E. Somssich (2000) Trends Plant Sci 5, 199-205.
Conserved amino acidsin WRKY proteins ofArabidopsis (red). Putative Zn-finger ligandsare highlighted in black. Eulgem et al. (2000) TIPS 5, 199-205.
The Plant Cell, Vol. 13, 1527–1539, July 2001. Evidence for an Important Role of WRKY DNA Binding Proteins in the Regulation of NPR1 Gene Expression Diqiu Yu, Chunhong Chen, and Zhixiang Chen 1 W-box: TTGAC
Binding of Arabidopsis proteinsto W-box sequence in NPR1’s promoter (PN1: 34 nt dsDNA). Npr1 upstream seq. -132 -99 Mutated Npr1 upstream seq. Antibodies to WRKYGQK peptideinhibited binding of both AtWRKY18and SA-induced binding activity to PN1 (Fig. 3). SA: - + - + - - AtWRKY18 AtWRKY18 Plant extract Pure protein Yu et al. (2001) Plant Cell 13, 1527-1539.
W-box is necessary for basal and SA-inducible NPR1 promoter activity. mRNA Reporter constructs contain 2419 ntfrom upstream of NPR1 ORF placedin front of GUS reporter. Used 25 indep. transformants/construct. Yu et al. (2001) Plant Cell 13, 1527-1539.
NPR1-independent NPR1-enhanced NPR1-enhanced NPR1-dependent SA-inducible WRKY genes. Yu et al. (2001) Plant Cell 13, 1527-1539.