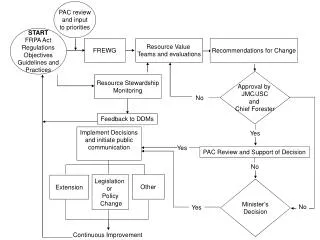
Identification of DNA Barcodes from Specific Phylum Using 16S rRNA Database Analysis
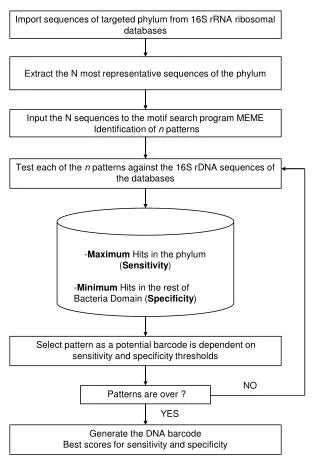
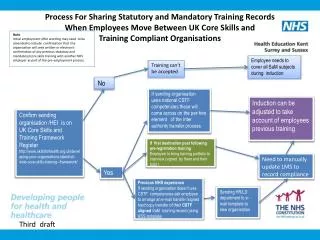
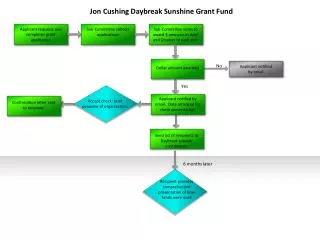
This study outlines a method for importing targeted phylum sequences from 16S rRNA ribosomal databases and extracting the most representative sequences. These sequences are then input into the MEME motif search program to identify n patterns. Each pattern is tested against 16S rDNA sequences to assess sensitivity (maximum hits in the target phylum) and specificity (minimum hits in other bacteria). Ultimately, patterns that meet the sensitivity and specificity thresholds can be considered potential barcodes, culminating in the generation of DNA barcodes to enhance microbial identification.

Identification of DNA Barcodes from Specific Phylum Using 16S rRNA Database Analysis
E N D
Presentation Transcript
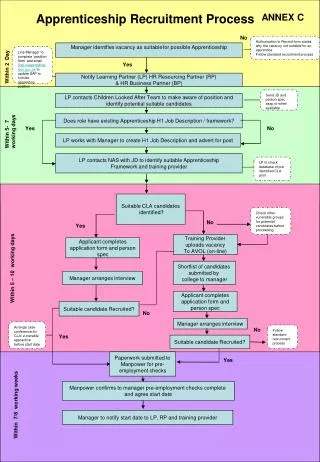
Import sequences of targeted phylum from 16S rRNA ribosomal databases Extract the N most representative sequences of the phylum Input the N sequences to the motif search program MEME Identification of n patterns Test each of the n patterns against the 16S rDNA sequences of the databases -Maximum Hits in the phylum (Sensitivity) -Minimum Hits in the rest of Bacteria Domain (Specificity) Select pattern as a potential barcode is dependent on sensitivity and specificity thresholds NO Patterns are over ? YES Generate the DNA barcode Best scores for sensitivity and specificity