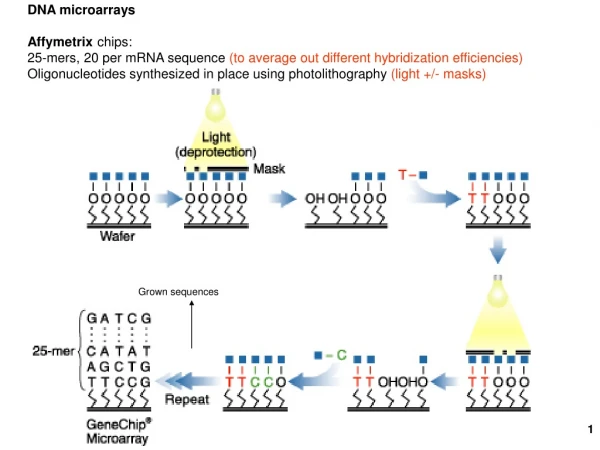
AFFYMETRIX chips
AFFYMETRIX chips. What are DNA chips/arrays?. Small silica (glass) chips covered with thousands of nucleotides of know sequence Underlying principle: Base pairing or hybridization Medium for matching known and unknown DNA samples Two types of arrays-Macro and Micro

AFFYMETRIX chips
E N D
Presentation Transcript
What are DNA chips/arrays? • Small silica (glass) chips covered with thousands of nucleotides of know sequence • Underlying principle: Base pairing or hybridization • Medium for matching known and unknown DNA samples • Two types of arrays-Macro and Micro Macro-Sample spot sizes >= 300 microns Micro-Sample spot sizes < 200 microns in diameter
Micro Arrays • Fabricated by high-speed robotics; generally on glass, but sometimes on nylon substrates • Known probes used to determine complementary binding • Enable massive parallel gene expression and gene studies • Dramatic increase in throughput • Two major applications: sequence identification and determination of gene expression level
Micro Arrays (contd.) • Two major variants, based on the property of the arrayed DNA sequence with known identity : 1) probe CDNA 2) AFFYMETRIX DNA chips
Micro Array: Another picture(intensity and color of each spot encode information on a specific gene from tested sample)
AFFYMETRIX • Pioneer in making tools for the genomic revolution • Applies semiconductor technology to life science • Manufactures AFFYMETRIX chips using a combination of photolithography and combinatorial chemistry
AFFYMETRIX chip: how is it made? • Very few synthesis steps • Produces arrays with hundreds of thousands of probes packed at very high density • Very high density helps obtain high-quality genomic data using small sample volumes • Number of synthesis steps determined by length of probes, not number
CONSTRUCTION • 5-inch square quartz wafer used • Wafer washed to ensure uniform hydroxylation across its surface • Quartz; good substrate for linker molecules, which are used to position the probes on the arrays • Wafer placed in bath of silane, which reacts with hydroxyl groups of quartz, and forms a matrix of covalently linked molecules
Construction (contd.) • Distance between silane molecules determines the probes’ packing density ( the arrays can hold over 500,000 probe locations) • Each of these probe locations harbors millions of identical DNA molecules. • Probe synthesis occurs in parallel: addition of an A,C,G or T nucleotide to multiple growing chains happens simultaneously
Probe synthesis • Photolithographic masks, carrying 18-20 square micron windows are placed over coated wafer. • Windows distributed over mask, depending on desired sequence of each probe • FIRST STEP of SYNTHESIS: UV light is shown over the mask, rendering the exposed linkers unprotected and available for nucleotide coupling. • CRITICAL: Alignment of mask with wafer before each synthesis step
Nucleotide attachment • Solution containing single type of deoxynucleotide (with removable protection group) is flushed over wafer’s surface • Nucleotide attaches to activated linkers, initiating synthesis • Although a very efficient process, some activated linkers fail to attach the new nucleotide: - Truncation used to prevent formation of probes with missing nucleotides
Nucleotide attachment (contd.) - Also, side chains of nucleotides are protected, to prevent the formation of branched oligonucleotides. • Another mask placed over wafer, for next round of deprotection and coupling • Algorithms used, to minimize mask usage • Process repeated, till probes reach their full length (typically 25 nucleotides)
Beyond synthesis • After synthesis is complete, the wafers are deprotected, diced and the resulting arrays are packaged in flowcell cartridges. • Depending on the number of probe locations per array, a single wafer can yield 49-400 arrays. • Quality control tests • Prepared chips well-suited to ambitious goals, like discovery of biological mechanisms and disease therapy
APPLICATIONS of Affymetrix chips TWO MAIN CLASSES: • Gene expression/RNA analysis • DNA analysis
Gene Expression • AFFYMETRIX developed a chip called Cytochrome P450, that could rapidly detect variation in gene activity affected by several therapeutics, including many beta-blockers and anti-depressants. • AFFYMETRIX teamed up with a bacterial genomics company, to develop an inexpensive chip to test drinking water for the presence of many different strains of pathogenic molecules.
Melanoma cells research • Researchers from Whitehead Institute and MIT identified a gene in melanoma cells that appears to play a crucial role in metastasis, the spread of cancer throughout the body. • Used data from Human Genome Project, DNA arrays • Screened the expression of about 10,000 genes • Found that a gene called rhoC plays a central role in metastasis
Dana-Farber Cancer Institute • Scientists from the institute have discovered a rare and lethal form of blood cancer that affects infants. • Data on next slide • MLL does not match with ALL or AML.
Colors representing gene activity (red is high, blue is low) show that MLL has a distinct pattern from both ALL and AML. Each vertical column of squares represents a tumor sample. Each horizontal row of squares represents the activity of one gene.
DNA analysis • AFFYMETRIX developed a chip to quickly scan a DNA sample for the presence of any of the mutant versions of the HIV virus. • A high-density DNA probe array, developed by AFFYMETRIX, has important applications for the identification of Mycobacterium tuberculosis species and the ability to detect of rifampicin resistance in a single platform.
References • http://www.gene-chips.com/GeneChips.html#What • http://www.gene-chips.com/sample1.html • http://www.polymerassemblytech.com/pages/4/ • http://www.ruderfinn.com/press_release.asp?release_id=79&bhcp=1 • http://www.affymetrix.com • http://www.dartmouth.edu/~cbbc/courses/bio4/bio4-1999/papers/JeffIsaacs.html • http://dnabridges.com/hottop/2000/2000NOV11.htm • http://www.genomenewsnetwork.org/articles/02_02/Whole_genome_leukemia_image1.shtml • http://pharmabiotech.ch/articles/20000701.htm • http://web.mit.edu/newsoffice/nr/2000/metastasis.html