Decoding Genes: Unraveling the Genetic Code and Protein Synthesis
330 likes | 398 Vues
Understand how DNA encodes amino acids, the central dogma, transcription and translation processes, and the differences between prokaryotic and eukaryotic cells. Discover the intricacies of genetic information flow.

Decoding Genes: Unraveling the Genetic Code and Protein Synthesis
E N D
Presentation Transcript
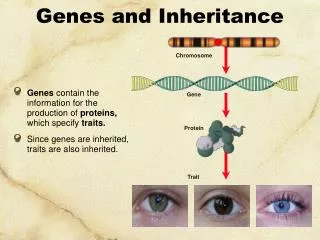
Genetic Code • How does the order of nucleotides in DNA encode information to specify the order of amino acids?
Genetic Code • Crick 1961 – elucidated the genetic code • Logic used - How many bases (nucleotides) are needed to code for 20 amino acids? • One base can code for 4 amino acids (41) • Two bases can code for 16 amino acids (42) • Three bases can code for 64 amino acids (43) • Therefore a sequence of three bases is the most reasonable number for a coden!
3 bases constitutes a codon (codes for an amino acid) with no space/markers between codons.
Codons and their amino acids • Nirenberg – used synthetic mRNA • Eg. UUUUUUU phenylalanine • Did not take long to determine amino acids and the corresponding 3 nucleotide sequence
Codon/amino acid relationship almost universal • e.g. Codon AGA arginine in Bacteria, Humans and all other organisms • except for Mitochondria and Chloroplasts and a few ciliates • What does this tell you?
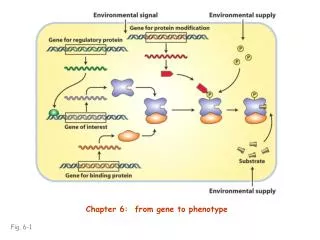
How does DNA make Proteins? • Central dogma: • DNA RNA ProteinTranscription Translation
RNA • Ribosomal RNA (rRNA) – made of several RNA molecules and over 50 proteins • Messenger RNA (mRNA) • Transfer RNA (tRNA)
Transcription (making mRNA) • Promotor – short sequence on DNA template strand where RNA polymerase binds. • Initiation – binding by RNA polymerase and starts unwinding DNA (17 base pairs long) • Elongation – 50 nucleotides added per second, no proof reading by RNA polymerase, therefore errors may occur. • Why is this not a big problem?
Transcription (cont’d) • Termination – stop sequences (series of GC forms a GC hairpin, slows down transcription. • Followed by 4 A which attaches 4 U, which are weak bonds, strand disassociation occurs
mRNA • mRNA now needs to travel out into cytoplasm • mRNA modified to prevent degradation by nucleases and phosphatases • Terminal 5’ end (usually A or G) is removed and is replaced with an unusual 5’-5’ linkage with GTP forming a 5’ cap. Protects end from degradation by nucleases and phosphotases. • 3’ end contain AAUAAA, poly A polymerase adds about 250 A’s to 3’ end long A tail. Needed to prevent degradation.
Translation • Making polypeptides
Advantage • In humans 1 to 1.5% of genome is exons • 24% are introns, rest of genome (75%) is non-incoding • Spliceosomes are large proteins that splice the exons together. • Human genes can be spliced together differently by spliceosomes. • Therefore 30,000 genes in humans can encode 120,000 different mRNA’s
Differences between Prokaryotes and Eukaryotes • Most eukaryotes posses Introns, Prokaryotes mostly do not! • Eukaryote mRNA contain transcripts of one gene. Prokaryote mRNA transcripts of several genes. • mRNA of eukaryotes must exit nucleus before translation can take place • Prokaryotes – translation starts at AUG codon Eukaryotic,start is also AUG, mRNA has a 5’ cap where translation is initiated. • Eukaryotic mRNA are modified, cap, tail and introns cut out • Eukaryotic rRNA are larger than those of Bacteria






![Getting Into Your Genes Another way to have a [healthy] baby!](https://cdn1.slideserve.com/2613166/getting-into-your-genes-another-way-to-have-a-healthy-baby-dt.jpg)