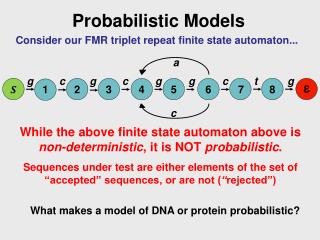
Probabilistic Graphical Models: Overview and Applications in Machine Learning
Explore the fundamentals of probabilistic graphical models, including Bayesian networks and Markov random fields. Learn about representation, conditional independence, inference techniques, and the role of graphical models in machine learning algorithms.

Probabilistic Graphical Models: Overview and Applications in Machine Learning
E N D
Presentation Transcript
Probabilistic Graphical Models COMP 790-90 Seminar Spring 2011
Outline • Introduction • Representation • Bayesian network • Conditional Independence • Inference: Variable elimination • Learning • Markov Random Field • Clique • Pair-wise MRF • Inference: Belief Propagation • Conclusion
Introduction • Graphical Model: + • Probability theory: ensures consistency, provides interface models to data. • Graph theory: intuitively appealing interface for humans, efficient general purpose algorithms. Probability Theory GraphTheory
Introduction • Modularity: a complex system is built by combining simpler parts. • Provides a natural tool for two problems: Uncertainty and Complexity • Plays an important role in the design and analysis of machine learning algorithms
Introduction • Many of the classical multivariate probabilistic systems are special cases of the general graphical model formalism: • Mixture models • Factor analysis • Hidden Markov Models • Kalman filters • The graphical model framework provides a way to view all of these systems as instances of common underlying formalism. • Techniques that have been developed in one field can be transferred to other fields • A framework for the design of new system
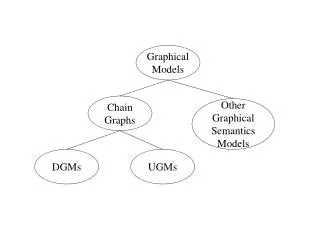
Representation • A graphical model represent probabilistic relationships between a set of random variables. • Variables are represented by nodes: • Binary events, Discrete variables, Continuous variables • Conditional (in)dependency is represented by (absence of) edges. • Directed Graphical Model: (Bayesian network) • Undirected Graphical Model: (Markov Random Field)
Outline • Introduction • Representation • Bayesian network • Conditional Independence • Inference: Variable elimination • Learning • Markov Random Field • Clique • Pair-wise MRF • Inference: Belief Propagation • Conclusion
Bayesian Network Parents • Directed acyclic graphs (DAG). • Directed edges give causality relationships between variables • For each variable X and parents pa(X) exists a conditional probability --P(X|pa(X)) • Discrete Variables: Conditional Probability Table(CPT) • Description of a noisy “causal” process
More Complex Example • Diagnose the engine start problem
More Complex Example • Computer-based Patient Case Simulation system (CPCS-PM) developed by Parker and Miller • 422 nodes and 867 arcs: 14 nodes describe diseases, 33 nodes describe history and risk factors, and the remaining 375 nodes describe various findings related to the diseases
Joint Distribution • P(X1,…Xn) • If the variables are binary, we need O(2n)parameters to describe P • For the wet grass example, need 2^4-1=15 parameters • Can we do better? • Key idea: use properties of independence.
Independent Random Variables • X is independent of Y iff for all values x,y • If X and Y are independent then • Unfortunately, most of random variables of interest are not independent of each other • The wet grass example
Conditional Independence • A more suitable notion is that of conditional independence. • X and Y are conditionally independent given Z • Notation: • The conditionally independent structure in the grass example • I(S,R|C) • I(C,W|S,R) C S R W
Parent Y1 Y2 X Descendent Y3 Non-descendent Y4 Conditional Independence • Directed Markov Property: Each random variable X, is conditionally independent of its non-descendents, given its parents Pa(X) • Formally,P(X|NonDesc(X), Pa(X))=P(X|Pa(X)) • Notation: I (X, NonDesc(X) | Pa(X))
Factorized Representation • Full Joint distribution is defined in terms of local conditional distributions(obtained via the chain rule) • Graphical Structure encodes conditional independences among random variables • Represent the full joint distribution over the variables more compactly • Complexity reduction • Joint probability of n binary variables O(2n) • Factorized form O(n*2k) • k: maximal number of parents of a node
Factorized Representation • The wetgrass example • P(C,S,R,W)=P(W|S,R)P(R|C)P(S|C)P(C) • Only need 1+2+2+4=9 parameters
Inference • Computation of the conditional probability distribution of one set of nodes, given a model and another set of nodes. • Bottom-up • Given Observation (leaves), the probabilities of the reasons can be calculated accordingly. • “diagnosis” from effects to reasons • Top-down • Knowledge influences the probability of the outcome • Predict the effects
Basic Computation y • The value of x depends on y • Dependency: conditional probability P(x|y) • Knowledge about y: prior probability P(y) • Product rule • Sum rule (Marginalization) • Bayesian rule x
Inference: Bottom UP Observe: wet grass (denoted by W=T) • Two possible causes: rain or sprinkle. Which is more likely? • Apply Bayes’ rule
Inference: Bottom UP Observe: wet grass (denoted by W=T) • Two possible causes: rain or sprinkle. Which is more likely? • Apply Bayes’ rule
Inference: Bottom UP Observe: wet grass (denoted by W=T) • Two possible causes: rain or sprinkle. Which is more likely? • Apply Bayes’ rule
Inference: Top-down • The probability that the grass will be wet given that it is cloudy. C S R W
Inference Algorithms • Exact inference problem in general graphical model is NP-hard • Exact Inference • Variable elimination • Message passing algorithm • Clustering and joint tree approach • Approximate Inference • Loopy belief propagation • Sampling (Monte Carlo) methods • Variational methods
Variable Elimination • Computing P(W=T) • Approach 1. Blind approach • Sum out all un-instantiated variables from the full joint • Computation Cost O(2n) • The wetgrass example • Number of additions: 14 • Number of products:? • Solution: explore the graph structure
Variable Elimination • Approach 2: Interleave sums and Products • The key idea is to push sums in as far as possible • In computation • First compute: • Then compute: • And so on • Computation Cost O(n*2k) • For wetgrass example • Number of Additions:? • Number of products:?
Learning • Learn parameters or structure from data • Structure learning: find correct connectivity between existing nodes • Parameter learning: find maximum likelihood estimates of parameters of each conditional probability distribution • A lot of knowledge (structures and probabilities) came from domain experts
Model Selection Method • Select a 'good' model from all possible models and use it as if it were the correct model • Having defined a scoring function, a search algorithm is then used to find a network structure that receives the highest score fitting the prior knowledge and data • Unfortunately, the number of DAG's on n variables is super-exponential in n. The usual approach is therefore to use local search algorithms (e.g., greedy hill climbing) to search through the space of graphs.
EM Algorithm • Expectation (E) step • Use current parameters to estimate the unobserved data • Maximization (M) step • Use estimated data to do ML/MAP estimation of the parameter • Repeat EM steps, until convergence
Outline • Introduction • Representation • Bayesian network • Conditional Independence • Inference • Learning • Markov Random Field • Clique • Pair-wise MRF • Inference: Belief Propagation • Conclusion
Markov Random Fields • Undirected edges simply give correlations between variables • The joint distribution is product of local functions over the cliques of the graph • where are the clique potentials, and Z is a normalization constant
The Clique • A clique • A set of variables which are the arguments of a local function • The order of a clique • The number of variables in the clique • Example: first order clique second order clique third order clique
φ1(i1) φ2(i2) φ3(i3) ψ12(i1, i2) ψ23(i2, i3) ψ14(i1, i4) ψ36(i3, i6) ψ25(i2, i5) φ4(i4) φ5(i5) φ6(i6) Observed image ψ45(i4, i5) ψ56(i5, i6) o1 o2 o3 i1 i2 i3 ψ69(i6, i9) ψ47(i4, i7) ψ58(i5, i8) φ7(i7) φ8(i8) φ9(i9) Underlying truth ψ78(i7, i8) ψ89(i8, i9) compatibility o4 o5 o6 i4 i5 i6 o7 o8 o9 i7 i8 i9 Pair-wise MRF • The order of cliques is at most two. • Commonly used in computer vision applications. • Infer underline unknown variables through local observation and the smooth prior
φ1(i1) φ2(i2) φ3(i3) ψ12(i1, i2) ψ23(i2, i3) ψ14(i1, i4) ψ36(i3, i6) ψ25(i2, i5) φ4(i4) φ5(i5) φ6(i6) Observed image ψ45(i4, i5) ψ56(i5, i6) o1 o2 o3 i1 i2 i3 ψ69(i6, i9) ψ47(i4, i7) ψ58(i5, i8) φ7(i7) φ8(i8) φ9(i9) Underlying truth ψ78(i7, i8) ψ89(i8, i9) o4 o5 o6 i4 i5 i6 o7 o8 o9 i7 i8 i9 Pair-wise MRF ψxy(ix, iy) is an nx* ny matrix. φx(ix) is a vector of length nx, where nx is the number of states of ix.
φ1(i1) φ2(i2) φ3(i3) ψ12(i1, i2) ψ23(i2, i3) ψ14(i1, i4) ψ36(i3, i6) ψ25(i2, i5) φ4(i4) φ5(i5) φ6(i6) Observed image ψ45(i4, i5) ψ56(i5, i6) o1 o2 o3 i1 i2 i3 ψ69(i6, i9) ψ47(i4, i7) ψ58(i5, i8) φ7(i7) φ8(i8) φ9(i9) Underlying truth ψ78(i7, i8) ψ89(i8, i9) o4 o5 o6 i4 i5 i6 o7 o8 o9 i7 i8 i9 Pair-wise MRF Given all the evidence nodes yi, we want to find the most likely state for all the hidden nodes xi, which is equivalent to maximizing
φ1(i1) φ2(i2) φ3(i3) ψ12(i1, i2) ψ23(i2, i3) ψ14(i1, i4) ψ36(i3, i6) ψ25(i2, i5) φ4(i4) φ5(i5) φ6(i6) Observed image ψ45(i4, i5) ψ56(i5, i6) o1 o2 o3 i1 i2 i3 ψ69(i6, i9) ψ47(i4, i7) ψ58(i5, i8) φ7(i7) φ8(i8) φ9(i9) Underlying truth ψ78(i7, i8) ψ89(i8, i9) o4 o5 o6 i4 i5 i6 o7 o8 o9 i7 i8 i9 Belief Propagation Beliefs are used to approximate this probability
Beliefs are used to approximate this probability Belief Propagation i2 o5 m2->5(i5) φ5(i5) i4 i5 i6 m6->5(i5) m4->5(i5) m8->5(i5) m8->5(x5) i8
Beliefs are used to approximate this probability m45(i5) i1 o4 m85(i5) m14(i4) φ4(i4) i4 m65(i5) m25(i5) m74(i4) i7 Belief Propagation i2 o5 ψ25(i2, i5) φ5(i5) ψ45(i4, i5) ψ56(i5, i6) x4 i5 i6 ψ58(i5, i8) i8
For every node ix Compute mzx(ix) for each neighbor iz Compute bx(ix) Belief Propagation φ(ix) and ψxy(ix,iy) N Does bx(ix) converge? Y Output most likely state for every node ix
Application: Learning Based Image Super Resolution • Extrapolate higher resolution images from low-resolution inputs. • The basic assumption: there are correlations between low frequency and high frequency information. • A node corresponds to an image patch • φx(xp): the probability of high frequency given observed low frequency • ψxy(xp, xq): the smooth prior between neighbor patches
Conclusion • A graphical representation of the probabilistic structure of a set of random variables, along with functions that can be used to derive the joint probability distribution. • Intuitive interface for modeling. • Modular: Useful tool for managing complexity. • Common formalism for many models.
References • Kevin Murphy, Introduction to Graphical Models, Technical Report, May 2001. • M. I. Jordan, Learning in Graphical Models, MIT Press, 1999. • Yijuan Lu, Introduction to Graphical Models, http:// www.cs.utsa.edu/~danlo/teaching/cs7123/Fall2005/Lyijuan.ppt. • Milos Hauskrecht, Probabilistic graphical models, http://www.cs.pitt.edu/~milos/courses/cs3710/Lectures/Class3.pdf. • P. Smyth, Belief networks, hidden Markov models, and Markov random fields: a unifying view, Pattern Recognition Letters, 1998.
References • F. R. Kschischang, B. J. Frey and H. A. Loeliger, 2001. Factor graphs and the sum-product algorithm IEEE Transactions on Information Theory, February, 2001. • Yedidia J.S., Freeman W.T. and Weiss Y, Understanding Belief Propagation and Its Generalizations, IJCAI 2001 Distinguished Lecture track. • William T. Freeman, Thouis R. Jones, and Egon C. Pasztor, Example-based super-resolution, IEEE Computer Graphics and Applications, March/April, 2002. • W. T. Freeman, E. C. Pasztor, O. T. Carmichael Learning Low-Level Vision International Journal of Computer Vision, 40(1), pp. 25-47, 2000.