Understanding Multiple Sequence Alignment in Evolutionary Biology
Learn how multiple sequence alignment (MSA) can reveal evolutionary forces shaping proteins. Explore MSA methods, consensus sequences, iterative alignment, and finding remote homologs using profile HMMs. Dive into a case study on the human kinome, discovering protein kinases and their diverse functions.

Understanding Multiple Sequence Alignment in Evolutionary Biology
E N D
Presentation Transcript
Multiple sequence alignment Lesson 4
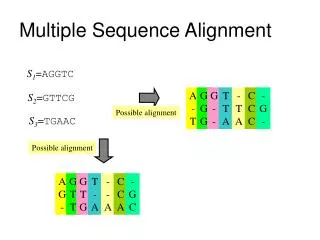
VTISCTGSSSNIGAG-NHVKWYQQLPG VTISCTGTSSNIGS--ITVNWYQQLPG LRLSCSSSGFIFSS--YAMYWVRQAPG LSLTCTVSGTSFDD--YYSTWVRQPPG PEVTCVVVDVSHEDPQVKFNWYVDG-- ATLVCLISDFYPGA--VTVAWKADS-- AALGCLVKDYFPEP--VTVSWNSG--- VSLTCLVKGFYPSD--IAVEWWSNG-- Like pairwise alignment BUT compare nsequences instead of 2 Each row represents an individual sequence Each column represents the ‘same’ position May be gaps in some sequences
MSA & Evolution MSA can give you a picture of the forces that shape evolution! • Important amino acids or nucleotides are not “allowed” to mutate • Less important positions change more easily
Conserved positions • Columns where all the sequences contain the same amino acids or nucleotides • Important for the function or structure VTISCTGSSSNIGAG-NHVKWYQQLPG VTISCTGSSSNIGS--ITVNWYQQLPG LRLSCTGSGFIFSS--YAMYWYQQAPG LSLTCTGSGTSFDD-QYYSTWYQQPPG
Consensus Sequence • A consensus sequence holds the most frequent character of the alignment at each column
Profile Profile = PSSM – Position Specific Score (probability) Matrix
Alignment methods There is no available optimal solution for MSA – all methods are heuristics: • Progressive/hierarchical alignment (Clustal) • Iterative alignment (mafft, muscle)
Progressive alignment A B C D E First step: Compute the pairwise alignments for all against all (6 pairwise alignments) the similarities are stored in a table
A B C D E Second step: • Cluster the sequences to create a tree (guide tree): • represents the order in which pairs of sequences are to be aligned • similar sequences are neighbors in the tree • distant sequences are distant from each other in the tree The guide tree is imprecise and is NOT the tree which truly describes the relationship between the sequences!
A B C D E Third step: sequence sequence sequence sequence 1. Align the most similar (neighboring) pairs
A B C D E Third step: sequence profile 2. Align pairs of pairs
Third step: profile sequence A B 3. Align out group C D E • Main disadvantages: • sub-optimal tree topology • Misalignments resulting from globally aligning a • pair of sequences will only cause further deterioration
Iterative alignment A B C DE Pairwise distance table Iterate until the MSA doesn’t change (convergence) Guide tree MSA A B C D E
Searching for remote homologs • Sometimes BLAST isn’t enough. • Large protein family, and BLAST only gives close members. We want more distant members • PSI-BLAST • Profile HMMs
Profile HMM • Similar to PSI-BLAST: also uses a profile • Takes into account: • Dependence among sites (if site n is conserved, it is likely that site n+1 is conserved part of a domain • The probability of a certain column in an alignment
PSI BLAST Vs. profile HMM PSI BLAST Profile HMM Less exact Faster More exact Slower
Case study: Using homology searching • The human kinome
Multi-tasking enzymes • Signal transduction • Metabolism • Transcription • Cell-cycle • Differentiation • Function of nervous and immune system • … • And more
How many kinases in the human genome? • 1950’s, discovery of that reversible phosphorylation regulates the activity of glycogen phosphorylase • 1970’s, advent of cloning and sequencing produced a speculation that the vertebrate genome encodes as many as 1001 kinases
How many kinases in the human genome? • 2001 – human genome sequence … • As well – databases of Genbank, Swissprot, and dbEST • How can we find out how many kinases are out there?
The human kinome • In 2002, Manning, Whyte, Martinez, Hunter and Sudarsanam set out to: • Search and cross-reference all these databases for all kinases • Characterize all found kinases
ePKs and aPKs Eukaryotic protein kinase (majority) catalytic domain Atypical protein kinases Sequence homology of the catalytic domain; additional regulatory domains are non-homologous No sequence homology to ePKs; some aPK subfamilies have structural similarity to ePKs
The search • Several profiles were built:based on the catalytic domain of: (a) 70 known ePKs from yeast, worm, fly, and human with >50% identity in the ePK domain (b) each subfamily of known aPKs • HMM-profile searches and PSI-BLAST searches were performed
The results… • 478 apKs • 40 ePKs • Total of 518 kinases in the human genome (half of the prediction in the 1970’s)
Classifying the kinases • Classification based on the catalytic domain • Classification based on the regulatory domains 189 sub-families of kinases
Comparison to other species • 209 subfamilies of ePKs in human, worm, yeast and fly
The human genome has x2 kinases (in number) as fly or worm. Many are aPKs. • Most of them are receptor tyrosine kinases (RTKs) The human-expanded kinase families function predominantly in processes of the: • Nervous system • Immune system • Angiogenesis • Hemopoiesis
The discovery of new kinases: a new front for battling human diseases
Correlating with human diseases • 160 kinases mapped to amplicons seen in tumors • 80 kinases mapped to amplicons in other major illnesses • Usually kinases are over-expressed in cancer and other diseases
Correlating with human diseases • 6 kinase inhibitors have been approved till today for the use against cancer • >70 other inhibitors are in clinical trials