5.2.3 Genomes and Gene Technologies
190 likes | 343 Vues
5.2.3 Genomes and Gene Technologies. outline the steps involved in sequencing the genome of an organism; outline how gene sequencing allows for genome-wide comparisons between individuals and between species (HSW7b); . Understanding DNA.

5.2.3 Genomes and Gene Technologies
E N D
Presentation Transcript
5.2.3 Genomes and Gene Technologies • outline the steps involved in sequencing the genome of an organism; • outline how gene sequencing allows for genome-wide comparisons between individuals and between species (HSW7b);
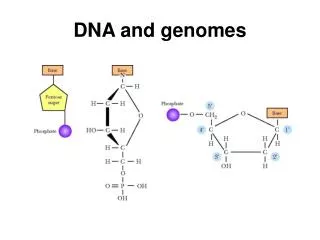
Understanding DNA Since discovering the structure of DNA, we can now use it for: • DNA profiling (genetic fingerprinting) in crime • Genomic sequencing to find the function of genes • Genetic engineering to make chemicals, GM organisms and for xenotransplantation • Gene therapy for treatment of diseases
Junk DNA • Genes code for production of polypeptides and proteins • This coding DNA is only 1.5% of the whole genome • The rest is non-coding or ‘junk’ DNA • We still don’t know what this ‘junk’ DNA does and research is ongoing • Genomics= the study of genomes and the ‘mapping’ (finding out the role of each gene) of organisms’ whole genome
Sequencing DNA • Can only sequence 750 base pairs at a time • Genome must be broken up into sections • Done a number of times with overlapping pieces • Overlapping sections analysed and put back together
Sequencing DNA • Genomes mapped to see where they have come from • Microsatellites (3-4 repetitive bases) used to help • Genome is broken into 100,000 base pairs (shotgun approach) • Sections put into BACS (bacterial artificial chromosomes and inserted into E-Coli • E-Coli grows and divides making clone libraries of the DNA
Sequencing BAC sections • Cells containing BACS are taken and grown • Restriction enzymes cut it into smaller fragments • Fragments separated by electrophoresis • Fragments sequenced using computer programmes that compare overlapping regions and put them back together The whole point of this is to get the entire base sequence of DNA of every organisms on the planet onto computer Then to figure out what each gene section does by comparing organisms DNA
Comparing Genomes • Comparative gene mapping = comparing genes for the same proteins across a range of organisms Why compare DNA? • clues to the importance of certain genes • shows evolutionary relationships • test the effect of mutations • identify genes that cause disease • mutant alleles can be revealed that make people more prone to heart disease etc.
DNA Separation • Electrophoresis: separates DNA fragments based on size • Restriction enzymes fragment DNA • DNA placed in wells at negative end • Immersed in buffer • Electric current switched on • DNA negatively charged and attracted to positive electrode • Shorter lengths move faster so move further • Dye then stains DNA to see where fragments have moved
In this diagram, the negative end is at the top and the positive electrodes are at the bottom so the DNA will move downwards... The shorter the fragments, the further down they will travel Sometimes the equipment is set up the other way with the positive electrode at the top
Southern Blotting • Fragments lifted from gel for analysis • Nylon sheet placed over gel with paper towels to blot it and left overnight • DNA fragments transferred to sheet for analysis • DNA labelled with radioactive marker • Photographic film shows DNA samples on finished sheet • This is called southern blotting after Edwin Southern who invented it • Radioactive probes can be used if you are looking for just one gene
DNA Probes • Short single stranded piece of DNA (50-80 bases long) • Complementary to the gene you are looking for • E.g. you know that the heart disease gene is AATTGCG you would create a strand complimentary to this and make it radioactive by replacing the phosphate in the nucleotides with a radioactive one e.g. 32P • You then expose the DNA strand to photographic film and find your DNA section • You could also use a fluorescent marker that emits colour when exposed to UV light • Copies of the probe can be added to any sample of DNA fragments and they will bind to any complementary base sequence as they are single stranded. This base pairing is called annealing
Why Use Probes? • Locate a gene for genetic engineering • Identify a gene on genomes from different species • Identify allele for diseases Disease Diagnosis A bit like immobilised enzymes, scientists can put probes on a fixed surface and apply the DNA. The DNA fragments that match will anneal to the fixed probes The DNA must first be fragmented and may be replicated using PCR... (polymerase chain reaction= artificial DNA replication)
Murder! • You are investigating a crime scene • You search the murder scene and find a skin cell on a knife that was used to stab the victim • As the murder happened a few months ago the DNA inside the cell is damaged and there are only a few base sequences of DNA • What do you do?
Polymerase Chain Reaction • Artificial DNA replication • Tiny samples of DNA replicated many times • Useful in forensics when you need many samples • Crime scene DNA can be multiplied (known as amplified) to get enough material for genetic profiling • Luckily this works because DNA is made of two backbones, is made of strands that run anti-parallel from 3’ end to 5’ end and are complimentary to one another
Limitations • Not identical to DNA replication • Can only replicate short sequences, not entire chromosoemes • Addition of primer molecules is necessary before starting • Cycles of heating and cooling are needed to separate and bind strands
How does PCR work? • DNA sample mixed with DNA nucleotides and DNA polymerase enzyme • Heated to 95⁰C breaking hydrogen bonds to make sample single stranded • Short lengths of single stranded DNA added (called primers) • Temperature reduced to 55⁰C allowing primers to bind (H bonds) and form small double stranded DNA sections • DNA polymerase binds to these strands • Temperature raised to 72⁰C for DNA polymerase to work by adding free nucleotides to the DNA • When DNA polymerase reaches the end it generates a new molecule of the double stranded DNA • This is repeated many times until many copies are produced • DNA polymerase is thermophillic as it is not denatured in extreme temperatures- it comes from a thermophillic bacteria which grows in hot springs up to 90⁰C
Automated ‘computer’ Sequencing • Rapid increase in sequencing time • Reaction mixture contains DNA polymerase • Many copies of single stranded template DNA fragment • Free nucleotides have fluorescent marker and are modified • If any of these nucleotides are added, DNA polymerase is ‘thrown off’ and the growing strand stops
How does a machine code DNA? • The primer DNA anneals (joins) to the end of the template strand • DNA polymerase attaches ready to build more DNA • DNA polymerase adds free nucleotides (same as normal DNA replication and PCR) • Many molecules of DNA made • When one of the labelled nucleotides is added, the reaction stops • Strands run through a machine with a laser and the machine records the labelled nucleotides What’s really going on? If you have a sequence of a known nucleotide and 6 unknowns e.g. A?????? And you label a C nucleotide with a colour, lets say blue You will get the DNA strand pairing A?????? T???C What is the unknown nucleotide? What about now? A?????? A????T