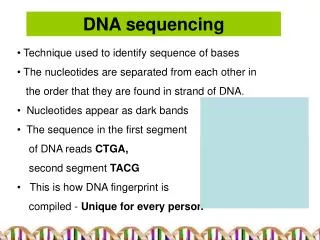
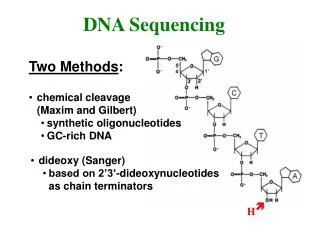
DNA Sequencing
This workshop presentation by Dr. Kabi R. Neupane covers the fundamentals of DNA sequencing, highlighting key techniques such as the RT-PCR product insertion into the pCR4-TOPO vector and the Sanger sequencing method. It delves into the use of DNA polymerase and primers like T3 and T7 in sequencing. Key topics include dideoxy nucleotides, chain termination, and how polyacrylamide gel electrophoresis separates DNA fragments. Attendees will learn about DNA sequencing applications in research and practical considerations for sample preparation.

DNA Sequencing
E N D
Presentation Transcript
0 DNA Sequencing Kabi R. Neupane, Ph.D. Leeward Community College ABE Workshop 2006
0 What to Sequence? • The RT-PCR product has been inserted into the pCR4-TOPO vector • Our goal • -Sequence the insert DNA
0 We Supplied Template DNA: Your plasmid Primer DNA: T3 or T7
0 Frederick Sanger • Discovered DNA sequencing by chain termination method • Nobel Prize 1 (1958) • Complete amino acid sequence of insulin • Nobel Prize 2 (1980) • For DNA sequencing
0 • DNA Sequencing exploits the DNA polymerase activity for deciphering DNA sequence • Modern DNA sequence use PCR technology in sequencing DNA Polymerase Action T3 Primer ATTAACCC TCACTAAAGG DNA Polymerase GACTAGTCCT GCAGGTTTAA AGGAATTCGC CCTT
0 One primer PCR Reaction DNA Polymerase Primer dATP dGTP dCTP dGTP Nucleotides NEW STRAND Template DNA
0 Dideoxy Nucleotides • Lack an -OH group at the 3-carbon position • Cannot add another nucleoside at that position • Prevent further DNA synthesis
0 Dideoxy nucleotides • Incorporation of a dideoxynucleotide to growing DNA strand terminates its further extension • Are added in small proportion • dATP ddATP • dGTP ddGTP • dCTP ddCTP • dTTP ddTTP
Use of Fluorescent Dyes Flurophores
0 Chain Termination
0 All Possible Terminations
0 Polyacrylamide Gel Electrophoresis Separates fragments based on size
0 DNA Sequence Files
0 Good or Bad?
0 Do not forget the other strand GGG ATATCACTCA GCATAATTGTTAAGTGACC T7 Primer
GenBank The Basic Local Alignment Search Tool (BLAST) finds regions of local similarity between sequences. 0 http://www.ncbi.nlm.nih.gov/
Involves fragment assembly using computer algorithms 0 Shotgun Sequencing Contigs
GREENWOOD MOLECULAR BIOLOGY FACILITY UNIVERSITY OF HAWAII AT MANOA 3050 Maile Way, Gilmore Hall 411, Honolulu, HI 96822 Phone: (808) 956-6718 Fax: (808) 956-9589 E-mail: biotech@hawaii.edu DNA SEQUENCING FORM PRIMARY INVESTIGATOR: _________________________________ DATE: _________________ YOUR NAME: ____________________________________________ DEPARTMENT: __________ ADDRESS: _____________________________________________________________________ _____________________________________________________________________ PHONE: _______________ FAX: ______________ E-MAIL: _____________________________ PURCHASE ORDER/REQUISITION NUMBER: __________________________________________ BILLING ADDRESS: _______________________________________________________________ <><><><><><><><><><><><><><><><><><><><><><><><><><><><><><><><><><><><><><><><><><><><><><><> Templates and primers should be supplied in ultrapure deionized water. Do not use Tris or other buffers. Please supply 12 µl of sample per reaction (template + one primer) in 0.5mL thin-walled PCR microcentrifuge tubes. Pre-reacted samples should be provided dry in 1.5mL centrifuge tubes. Please do not attach tags to tubes. Samples may not be run simultaneously. PCR productsPlasmidSS TemplatesCosmid TEMPLATE 20ng/100 bp 0.5-1.0 μg 0.25-0.5 μg 0.5-1.0 μg PRIMER 3.2 pmole 3.2 pmole 0.8 pmole 25pmole <><><><><><><><><><><><><><><><><><><><><><><><><><><><><><><><><><><><><><><><><><><><><><><> SAMPLE NAME: 1. _______________ 11. _______________ 21. _______________ 2. _______________ 12. _______________ 22. _______________ 3. _______________ 13. _______________ 23. _______________ 4. _______________ 14. _______________ 24. _______________ 5. _______________ 15. _______________ 25. _______________ 6. _______________ 16. _______________ 26. _______________ 7. _______________ 17. _______________ 27. _______________ 8. _______________ 18. _______________ 28. _______________ 9. _______________ 19. _______________ 29. _______________ 10. _______________ 20. _______________ 30. _______________ SPECIAL INSTRUCTIONS: __________________________________________________________ DATA DELIVERY: 3.5’’ disk or ZIP disk (must provide) ______ FTP _______ E-mail attachment _______ Electrophoregram print-out ($2.00 per sample) _________ Type of Computer used: MAC _______ PC _________ SIGNATURE: _________________________________________