Co-regulation Identification Using Probabilistic Relational Models
This research by Christoforos Anagnostopoulos, supervised by Dirk Husmeier, employs Probabilistic Relational Models (PRMs) for identifying co-regulation in gene expression data. The study integrates disparate data sources, including problematic promoter sequences and mRNA expression data from overexpressed genes. It introduces a Bayesian modeling framework to analyze transcriptional modules, evaluate gene interaction, and assess motif presence. The presentation aims to outline the Segal model's structures, criticisms, and the implications of PRMs for future biological research, particularly in gene regulation and network modeling.

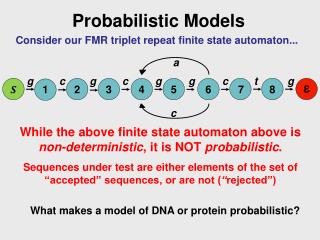
Co-regulation Identification Using Probabilistic Relational Models
E N D
Presentation Transcript
Identifying co-regulation using Probabilistic Relational Models by Christoforos Anagnostopoulos BA Mathematics, Cambridge University MSc Informatics, Edinburgh University supervised by Dirk Husmeier
General Problematic Promoter sequence data ...ACGTTAAGCCAT... ...GGCATGAATCCC... Bringing together disparate data sources:
General Problematic Promoter sequence data ...ACGTTAAGCCAT... ...GGCATGAATCCC... Bringing together disparate data sources: Gene expression data gene 1: overexpressed gene 2: overexpressed ... mRNA
General Problematic Promoter sequence data ...ACGTTAAGCCAT... ...GGCATGAATCCC... Bringing together disparate data sources: Gene expression data gene 1: overexpressed gene 2: overexpressed ... mRNA Protein interaction data protein 1 protein 2 ORF 1 ORF 2 -------------------------------------------------- AAC1 TIM10 YMR056C YHR005CA AAD6 YNL201C YFL056C YNL201C Proteins
Our data Promoter sequence data ...ACGTTAAGCCAT... ...GGCATGAATCCC... Gene expression data gene 1: overexpressed gene 2: overexpressed ... mRNA
Bayesian Modelling Framework Bayesian Networks
Bayesian Modelling Framework Conditional Independence Assumptions Factorisation of the Joint Probability Distribution Bayesian Networks UNIFIED TRAINING
Bayesian Modelling Framework Probabilistic Relational Models Bayesian Networks
Aims for this presentation: Briefly present the Segal model and the main criticisms offered in the thesis Briefly introduce PRMs Outline directions for future work
The Segal Model Module 1 Module 2 Cluster genes into transcriptional modules... ? gene
The Segal Model Module 1 Module 2 P(M = 1) P(M = 2) gene
The Segal Model Module 1 How to determine P(M = 1)? P(M = 1) gene
The Segal Model Motif Profile motif 3: active motif 4: very active motif 16: very active motif 29: slightly active Module 1 How to determine P(M = 1)? gene
The Segal Model Predicted Expression Levels Array 1: overexpressed Array 2: overexpressed Array 3: underexpressed ... Motif Profile motif 3: active motif 4: very active motif 16: very active motif 29: slightly active Module 1 How to determine P(M = 1)? gene
The Segal Model Predicted Expression Levels Array 1: overexpressed Array 2: overexpressed Array 3: underexpressed ... Motif Profile motif 3: active motif 4: very active motif 16: very active motif 29: slightly active Module 1 How to determine P(M = 1)? P(M = 1) gene
The Segal model PROMOTER SEQUENCE
The Segal model PROMOTER SEQUENCE MOTIF PRESENCE
The Segal model PROMOTER SEQUENCE MOTIF MODEL MOTIF PRESENCE
The Segal model MOTIF PRESENCE MODULE ASSIGNMENT
The Segal model MOTIF PRESENCE REGULATION MODEL MODULE ASSIGNMENT
The Segal model MODULE ASSIGNMENT EXPRESSION DATA
The Segal model MODULE ASSIGNMENT EXPRESSION MODEL EXPRESSION DATA
Learning via hard EM HIDDEN
Learning via hard EM Initialise hidden variables
Learning via hard EM Initialise hidden variables Set parameters to Maximum Likelihood
Learning via hard EM Initialise hidden variables Set parameters to Maximum Likelihood Set hidden values to their most probable value given the parameters (hard EM)
Learning via hard EM Initialise hidden variables Set parameters to Maximum Likelihood Set hidden values to their most probable value given the parameters (hard EM)
Motif Model OBJECTIVE: Learn motif so as to discriminate between genes for which the Regulation variable is “on” and genes for which it is “off”. r = 1 r = 0
Motif Model – scoring scheme high score: ...CATTCC... low score: ...TGACAA...
Motif Model – scoring scheme high score: ...CATTCC... low score: ...TGACAA... high scoring subsequences ...AGTCCATTCCGCCTCAAG...
Motif Model – scoring scheme high score: ...CATTCC... low score: ...TGACAA... high scoring subsequences ...AGTCCATTCCGCCTCAAG... low scoring (background) subsequences
Motif Model – scoring scheme high score: ...CATTCC... low score: ...TGACAA... high scoring subsequences ...AGTCCATTCCGCCTCAAG... promoter sequence scoring low scoring (background) subsequences
Motif Model SCORING SCHEME P ( g.r = true | g.S, w ) parameter set w: can be taken to represent motifs
Motif Model SCORING SCHEME P ( g.r = true | g.S, w ) parameter set w: can be taken to represent motifs Maximum Likelihood setting Most discriminatory motif
Motif Model – overfitting TRUE PSSM
Motif Model – overfitting typical motif: ...TTT.CATTCC... TRUE PSSM high score
Motif Model – overfitting typical motif: ...TTT.CATTCC... TRUE PSSM high score INFERRED PSSM Can triple the score!
Regulation Model For each module m and each motif i, we estimate the association umi P ( g.M = m | g. R ) is proportional to
Regulation Model: Geometrical Interpretation The(umi )i define separating hyperplanes Classification criterion is the inner product: Each datapoint is given the label of the hyperplane it is the furthest away from, on its positive side.
Regulation Model: Divergence and Overfitting pairwise linear separability overconfident classification Method A: dampen the parameters (eg Gaussian prior) Method B: make the dataset linearly inseparable by augmentation
Erroneous interpretation of the parameters Segal et al claim that: When umi = 0, motif iis inactive in module m When umi > 0 for all i,m, then only the presence of motifs is significant, not their absence
Erroneous interpretation of the parameters Segal et al claim that: When umi = 0, motif iis inactive in module m When umi > 0 for all i,m, then only the presence of motifs is significant, not their absence Contradict normalisation conditions!
Sparsity INFERRED PROCESS TRUE PROCESS
Sparsity Reconceptualise the problem: Sparsity can be understood as pruning Pruning can improve generalisation performance (deals with overfitting both by damping and by decreasing the degrees of freedom) Pruning ought not be seen as a combinatorial problem, but can be dealt with appropriate prior distributions
Sparsity: the Laplacian How to prune using a prior: choose a prior with a simple discontinuity at the origin, so that the penalty term does not vanish near the origin every time a parameter crosses the origin, establish whether it will escape the origin or is trapped in Brownian motion around it if trapped, force both its gradient and value to 0 and freeze it Can actively look for nearby zeros to accelerate pruning rate
Results: generalisationperformance Synthetic Dataset with 49 motifs, 20 modules and 1800 datapoints
Results: interpretability DEFAULT MODEL: LEARNT WEIGHTS TRUE MODULE STRUCTURE LAPLACIAN PRIOR MODEL: LEARNT WEIGHTS
Aims for this presentation: Briefly present the Segal model and the main criticisms offered in the thesis Briefly introduce PRMs Outline directions for future work
Probabilistic Relational Models How to model context – specific regulation? Need to cluster the experiments...