DNA Computing Computation with the Code of Life
DNA computing, introduced by Len Adleman in 1994, leverages the combinatorial properties of DNA to solve complex problems encoded as DNA strings. This groundbreaking approach combines molecular biology with computer science, offering high data density and the potential for massively parallel processing. The DNA strand’s minute size (0.35nm per base) allows for computations at a molecular level, presenting a revolutionary method with applications ranging from data storage to in-body computation. Despite challenges like error rates and complexity, the future of DNA computing remains promising.

DNA Computing Computation with the Code of Life
E N D
Presentation Transcript
DNA ComputingComputation with the Code of Life Michael Ang <m@michaelang.com> Interactive Telecommunications Program New York University February 16, 2007
What is it? • Invented by Len Adleman (1994) • Realized combinatorial properties of DNA could be used to solve problems encoded as strings of DNA
Why is it interesting? • Cross-discipline (CS meets Molecular Biology) • High data density • Data and computation happens at molecular level • Each base of DNA is 0.35nm • Massively parallel • 1012 or more copies of DNA in a test tube • Use biological enzymes to make these copies • Energy efficient • Potential to perform computation inside the body • Though big challenges before a reality
What is DNA? • Source code to life • Instructions for building and regulating cells • Data store for genetic inheritance • Cellular machinery (enzymes) translates DNA into proteins, duplicates, repairs, etc. • Think of enzymes as hardware, DNA as software
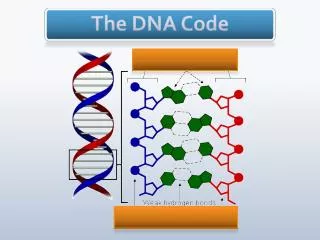
What is DNA made of? • Composed of four nucleotides (+ sugar-phosphate backbone) • A – Adenine • T –Thymine • C – Cytosine • G – Guanine • Bond in pairs • A – T • C – G
How does it work? • Use specially coded DNA as initial conditions for biological reaction • Natural enzymes duplicate DNA • Matching DNA base pairs attach to each other • Find answer in resulting soup of DNA strands
Travelling Salesman • Simple problem (at small scale) • Complexity scales exponentially
Algorithm • Generate all possible routes • Select routes that start with the initial city and end with the destination city • Select itineraries with the correct number of cities • Select itineraries that contain each city once
Algorithm • Generate all possible routes • Select routes that start with the initial city and end with the destination city • Select itineraries with the correct number of cities • Select itineraries that contain each city once
Annealing • Use “a lot” (1013) copies of each city and path
Algorithm • Generate all possible routes • Select routes that start with the initial city and end with the destination city • Select itineraries with the correct number of cities • Select itineraries that contain each city once
Algorithm • Generate all possible routes • Select routes that start with the initial city and end with the destination city • Select itineraries with the correct number of cities • Select itineraries that contain each city once
Gel electrophoresis • “Race” the DNA through a gel • Shorter lengths go faster
Algorithm • Generate all possible routes • Select routes that start with the initial city and end with the destination city • Select itineraries with the correct number of cities • Select itineraries that contain each city once
Affinity purify • Pull out strands matching one city at a time
We have our answer • Sequence result or perform more PCR reactions to determine combinatorically • Took Adleman 7 days to run through the steps • Actual “computation” took minutes
Problems/Challenges • Relatively high error rate in DNA • Sometimes bases don’t align properly • Problem complexity still scales exponentially • 200 city problem might take DNA weighing more than Earth • Many people doubtful…
Can I do it? • If you have access to a lab, yes • DIY seems possible (<$500) Personal Biocomputing – Eugene Thacker
Further Research • Use to break DES (1995) • Turing Machine (at least… “similar to”) (2001) • Play tic-tac-toe (2003) • DNA computer to detect and treat cancer (2004) • Use enzymes and DNA to create state machine using mRNA as input • Output DNA sequence to suppress disease causing gene • In vitro (petri dish) so far • Much harder in vivo (living cell)
A nanoscale programmable computing machine with input, output, software and hardware made of biomolecules Nature 414, 430-434 (2001) • 1012 automata run independently and in parallel • on potentially distinct inputs • in 120 ml • at room temperature • at combined rate of 109 transitions per second • with accuracy greater than 99.8% per transition, • consuming less than 10-10 Watt.
So what? • Computation at molecular level • Computation (potentially) within cells • Operate with cellular messages as input and output • Operate near theoretical power limits • Computer Science implemented in Biology • Practical applications still a dream…
To correctly gauge the practicality of molecular computing will require inputs from experts in a wide variety of fields including: biology, chemistry, computer science, engineering, mathematics and physics. - Len Adleman (1995)
References • “What is DNA” - Genetics Home Reference • “DNA Computer Could Target Cancer” – Nanotech Web • “DNA Computing: A Primer” – Arstechnica • “On Constructing a Molecular Computer” – Len Adleman • “Breaking DES using a molecular computer” - D. Boneh, C. Dunworth, and R. Lipton • “Injectable Medibots: Programmable DNA could diagnose and treat cancer” - Alexandra Goho • “First game-playing DNA computer revealed” – New Scientist • “Programmable and autonomous computing machine made of biomolecules” - Benenson, Paz-Elizur, Adar, Keinan, Livneh & Shapiro • “Personal Biocomputing” – Eugene Thacker • “Biological Nanocomputer” – Weizman Institute • “Computer Made from DNA and Enzymes” – National Geographic News
Thanks Michael Ang <m@michaelang.com> Interactive Telecommunications Program New York University February 16, 2007