DNA sequencing
E N D
Presentation Transcript
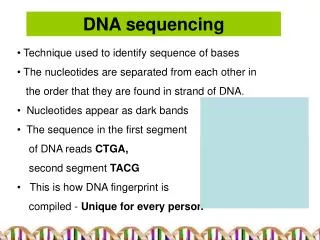
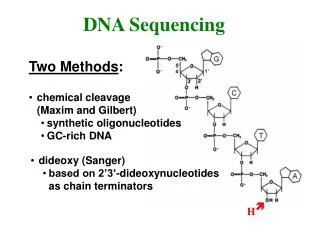
The first generation The Sanger dideoxy method of DNA sequencing
This method was costly, at around $10 per base pair in 1985, but the development of automated sequencing systems and advancements in technology reduced the price to $1 per base by 1995, and allowed sequencing of up to 100,000 bases per day. • The cost dropped further to $0.10 per base in 1998 with the development of the ABI Prism sequencer, which made it possible toundertake larger-scale sequencing projects. • In the context of highthroughput shotgun genomic sequencing, Sanger sequencing costs on the order of $0.50 per kilobase.
In search of the $1000 genome • National Institutes of Health (NIH), specifically the NIH’s National Human Genome Research Institute (NHGRI), has actively sponsored researchers with $99.5 million of external funding to develop sequencing technologies since the completion of the Human Genome Project. • $100,000 initially and then as little as $1000.
The second generation: sequencing-by-synthesis • Pyrosequencing ($1.5million, 2008) • Illumina ($48,000 ?, 2010) • SOLiD
control Enzyme mix 5ul Substrare mixture 5ul 100pmol primer 1ul 500fmol template 5ul 10xBuffer 10ul 水 74ul GCAGGATG 間隔2min dNTP 0.2x S E A G T C
The third generation:direct measuremeant or synthesis-free • Nanopore