Microarray
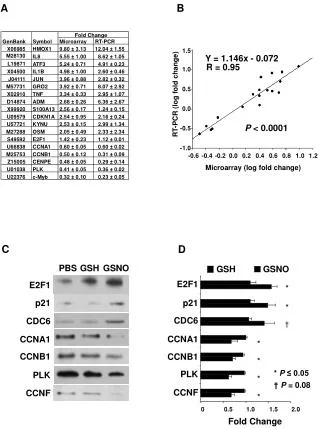
Materials & Methods. Microarray oligonucleotide microarrays [Hu6800 and Hu35KsubA GeneChips (Affymetrix, Santa Clara, CA)] containing a total of 16,063 probe sets representing 14,030 GenBank and 475 The Institute for Genomic Research (TIGR) accession. Supervised learning method.


Microarray
E N D
Presentation Transcript
Materials & Methods Microarray oligonucleotide microarrays [Hu6800 and Hu35KsubA GeneChips (Affymetrix, Santa Clara, CA)] containing a total of 16,063 probe sets representing 14,030 GenBank and 475 The Institute for Genomic Research (TIGR) accession Supervised learning method Of 314 tumor samples and 98 normal tissue samples processed, 218 tumors and 90 normal tissue samples passed quality control criteria and were used for subsequent data analysis. The resulting dataset contained ~5 million gene expression values.
14 tumor classes: BR, breast adenocarcinoma; PR, prostate adenocarcinoma; LU, lung adenocarcinoma; CR, colorectal adenocarcinoma; LY, lymphoma; BL, bladder transitional cell carcinoma; ML, melanoma; UT, uterine adenocarcinoma; LE, leukemia; RE, renal cell carcinoma; PA, pancreatic adenocarcinoma; OV, ovarian adenocarcinoma; ME, pleural mesothelioma; CNS, central nervous system hyperplane : best separates samples from two classes
(NKI) • analysis on primary breast tumors of 117 young patients • - 34 : developed distant metastases within 5 years • - 44 : disease-free after a period of at least 5 years • - 18 : BRCA1 germline mutations • - 2 : BRCA2 carriers • the signatures of breast cancer gene expression reported to date allow • for patient-tailored therapy strategies
If such results are confirmed by others, doctors may one day be able to discern which patients need the most intensive therapy based in part on how closely their expression profiles match a standard good or poor prognosis profile.
Agendia's first product is MammaPrint(R), a gene expression profiling service to assess the recurrence risk in breast cancer patients. MammaPrint(R) provides valuable assistance for oncologists in formulating tailor-made treatment plans. 檢測費用:US$2196
Rapid Communications Broadly Altered Gene Expression in Blood Leukocytes in Essential Hypertension Is Absent During Treatment (Hypertension. 2004;43:947-951.) Helena Chon, Carlo A.J.M. Gaillard, Brenda B. van der Meijden, Hilde M. Dijstelbloem, Rob J. Kraaijenhagen, Dik van Leenen, Frank C.P. Holstege, Jaap A. Joles, Hans A.R. Bluyssen, Hein A. Koomans, Branko Braam From the Department of Nephrology (H.C., J.A.J., H.A.R.B., H.A.K., B.B.), University Medical Center, Utrecht, The Netherlands; Department of Internal Medicine (C.A.J.M.G.) and Department of Clinical Chemistry (B.B.v.d.M., H.M.D., R.J.K.), Meander Medical Center, Amersfoort, The Netherlands; and Genomics Laboratory (D.v.L., F.C.P.H.), University Medical Center, Utrecht, The Netherlands.
Drugs used were angiotensin-converting enzyme (ACE) inhibitors (n=3), b-blockers (n=2), or a combination of b-blockade and diuretic (n=1).
treated untreated • In untreated hypertension patients, 680 genes were differentially regulated (314 up and 366 down). • In the treated patients, these changes were virtually absent (4 genes up, 3 genes down).
oxidative stress-related genes • inflammation-related genes • Blood pressure regulation genes