Protein Purification
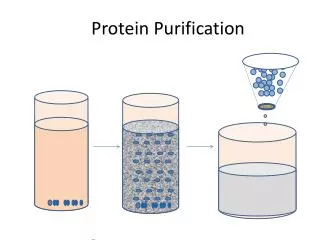
Protein Purification. Tutorial Minnie Murugesan Dr. Scott’s Group 07-12-05. Eventful Protein Purification. Grow cells in media (vector+tag) Centrifuge, Collect the pellet Lyse the cells (appropriate buffer). Pilot Expression SDS PAGE, Assay. Purification Strategy. Characterization

Protein Purification
E N D
Presentation Transcript
Protein Purification Tutorial Minnie Murugesan Dr. Scott’s Group 07-12-05
Eventful Protein Purification • Grow cells in media (vector+tag) • Centrifuge, Collect the pellet • Lyse the cells (appropriate buffer) Pilot Expression SDS PAGE, Assay Purification Strategy Characterization Mass Spectroscopy X-ray Crystallography Functional Assay Solubility Aggregation Recombination
OUTLINE • Chromatography techniques • Affinity Chromatography (AC) • Hydrophobic Interaction Chromatography (HIC) • Ion Exchange Chromatography (IEC) • Gel Filtration (GF) • Capillary electrochromatography (CEC) • Other New Strategies for Protein Purification • Solubility, Aggregation and Re-folding of Proteins Invented by a Russian botanist named Mikhail Tswett in 1903. He separated plant pigments using glass columns packed with calcium carbonate.
Protein Purification Strategies Evaluate an assay for the protein of interest Shortlist a method to have a reasonable source for that activity (http://www5.amershambiosciences.com/aptrix/upp00919.nsf/Content/LabSep_EduC%5CAboutPurBiom%5CHowToCombine)
For Membrane Proteins Three Phase Strategy (www.amershambiosciences.com)
Chromatographic Modes of Protein Purification (Christian G. Huber, Biopolymer Chromatography, Encylcopedia in analytical chemistry, 2000)
Affinity Chromatography Affinity Chromatography Surface bound with Epoxy, aldehyde or aryl ester groups Metal Interaction Chromatography Surface bound with Iminodiacetic acid + Ni2+/Zn2+/Co2+ (Christian G. Huber, Biopolymer Chromatography, Encylcopedia in analytical chemistry, 2000)
Metal Interaction Chromatography (AC) Points to Note: Avoid chelating agents Increasing incubation time 3. Slow gradient elution (www.qiagen.com)
Affinity Chromatography Binding Capacity (mg/ml) medium 12mg of histag proteins (MW= 27kDa) Depends on Molecular weight Degree of substitution /ml medium ~15mmol Ni2+ Backpressure ~43psi Change the guard column filter (Christian G. Huber, Biopolymer Chromatography, Encylcopedia in analytical chemistry, 2000)
Hydrophobic region Hydrophobic Interaction Chromatography Biopolymer (phenyl agarose - Binding Surface) Driving force for hydrophobic adsorption Water molecules surround the analyte and the binding surface. When a hydrophobic region of a biopolymer binds to the surface of a mildly hydrophobic stationary phase, hydrophilic water molecules are effectively released from the surrounding hydrophobic areas causing a thermodynamically favorable change in entropy. Temperature plays a strong role Ammonium sulfate, by virtue of its good salting-out properties and high solubility in water is used as an eluting buffer (Christian G. Huber, Biopolymer Chromatography, Encylcopedia in analytical chemistry, 2000)
Globular Protein Maintenance of conformation while interacting with tentacle ion exchanger Deformation due to interaction with conventional ion exchanger Ion Exchange Chromatography Fractogel matrix is a methacrylate resin upon which polyelectrolyte Chains (or tentacles) have been grafted. (Novagen) (www.novagen.com)
Gel Filtration (http://lsvl.la.asu.edu/resources/mamajis/chromatography/chromatography.html)
Immuno Affinity Chromatography (http://www.cellmigration.org/resource/discovery/discovery_proteomics_approaches.html)
DNA Binding Proteins ATP immobilized on polyacrylamide resin Heparin Sepharose Negatively charged proteins (pI >7) are not captured/separated effectively. (www.novagen.com)
Capillary Electrochromatography • CEC is an electrokinetic separation technique • Fused-silica capillaries packed with stationary phase • Separation based on electroosmotically driven flow • Higher selectivity due to the combination of chromatography and electrophoresis • Fused silica tube filled with porous methacrylamide-stearyl methacrylate-dimethyldiallyl ammonium chloride monolithic polymers, 80 x 0.5mm i.d., 5.5kV. High Plate count ~ 400,000 Height Equivalent to a Theoretical Plate /Plate Count (HETP) H = L/N number of plates N = 16(t/W)2 where L = column length, t = retention time, and W = peak width at baseline (http://www.capital-hplc.co.uk)
CEC columns AC, IEC columns CEC column NP, RP columns
Commercially available protein purification kits GST•Bind™ Purification Kits His•Bind® Purification Kits Magnetight™ Oligo d(T) Beads MagPrep® Streptavidin Beads Protein A and Protein G Plus Agaroses S•Tag™ Purification Kits Streptavidin Agarose T7•Tag™ Affinity Purification Kit ProteoSpin™ CBED (Concentration, Buffer Exchange and Desalting) Maxi Kit — Effectively desalts and concentrates up to 8 mg of protein with an efficient, easy-to-use protocol.(Norgen Biotek Corporation) ProteoSpin™ Detergent Clean-up Micro Kit — Provides a fast and effective procedure to remove detergents including SDS, Triton® X-100, CHAPS, NP-40 and Tween 20. (http://www.emdbiosciences.com)
No air bubbles • (Priming) • Use degassed buffers Injector Module 2 1 Column Inlet 3 Detector 4 Fraction Collector 5 Fast Protein Liquid Chromatograph (FPLC) (www.pharmacia.com)
Protein Analysis (http://www.cellmigration.org/resource/discovery/discovery_proteomics_approaches.html)
Detection of Proteins by Derivatization with Higher Sensitivity 1000 times more sensitive than UV-Vis detection (Christian G. Huber, Encylcopedia in analytical chemistry, 2000)
Solubility of a protein • Depends strongly on the composition of the lysis buffer. • Salt concentration • Freeze-thaw protocol • * Freeze quickly on dry ice and leave for 3 min. • * Thaw immediately at 42 °C. Vortex vigorously to mix well. • * Repeat the two previous steps three more times (4 cycles in all). Membrane proteins 1. Removal of unbroken cells from the cell lysate by low speed centrifugation (20 min at 10,000 g). 2. Isolation of the membrane particles from the supernatant by ultracentrifugation (60 min at >100 000 g). 3. Washing of the membrane particle to remove all soluble proteins. Solubilization of protein from the membrane particles by a mild detergent.(detergent: protein ratio = 1:10) 5. Phosphate buffers(0.1M-0.5M), 5-50% glycerol helps. (http://www.ls.huji.ac.il/~purification)
Protein Aggregation • Numerous physicochemical stresses can induce protein aggregation: • Heat, pressure, pH, agitation, freeze-thawing, dehydration, heavy metals, • phenolic compounds, and denaturants. (http://www.integritybio.com/protein%20aggregation.htm)
Solubilization of Aggregated Proteins Denaturation and Renaturation Variables Good starting point Buffer composition (pH, ionic strength) 50 mM Tris-HCl, pH 7.5 Incubation temperature 30°C Incubation time 60 min Concentration of solubilzing agent 6 M guanidine-HCl or 8 M urea Total protein concentration 1-2 mg/ml Re-folding of Proteins The addition of a mixture of reduced and oxidized forms of low molecular weight thiol reagent usually provides the appropriate redox potential to allow formation and reshuffling of disulfide bonds (1-3 mM reduced thiol and a 5:1 to 1:1 ratio of reduced to oxidixed thiol) The most commonly used are glutathione, cysteine and cysteamine. (www.biovectra.com)
Reagents used for Re-folding of proteins (http://www.ls.huji.ac.il/~purification)
Reagents used for Re-folding of proteins (Continued) (http://www.ls.huji.ac.il/~purification)
6xHis Tagged Protein Detection Directly on the Gel (from Pierce) E. coli lysates expressing 6xHis-tagged proteins, stained with the Pierce 6xHis Protein Tag Staining Kit (www.piercenet.com)
University of Oklahoma • Recombinant Protein Solubility Prediction • Type (or cut and paste) your protein sequence below, click on the "Submit" button, and the solubility probability of your protein will be calculated. • The statistical model predicts protein solubility assuming the protein is being over-expressed in Escherichia coli. • The input protein sequence has a 73.4 percent chance of insolubility when overexpressed in E. coli. - mbh8 NADH dehydrogenase subunit - integral membrane protein (NP_579159) • The input protein sequence has a 80.5 percent chance of solubility when overexpressed in E. coli. - replication factor A related protein (NP_579749) (www.biotech.ou.edu)