Hidden Markov Models
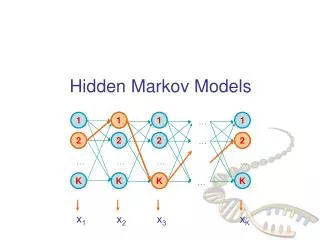
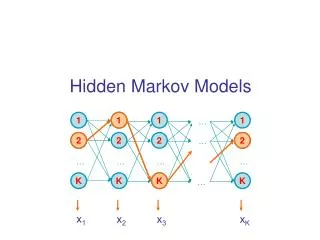
Hidden Markov Models. A first-order Hidden Markov Model is completely defined by: A set of states. An alphabet of symbols. A transition probability matrix T=(t ij ) An emission probability matrix E=(e iX ). Linear Architecture. Loop Architecture. Wheel Architecture. Basic Ideas.

Hidden Markov Models
E N D
Presentation Transcript
Hidden Markov Models A first-order Hidden Markov Model is completely defined by: • A set of states. • An alphabet of symbols. • A transition probability matrix T=(tij) • An emission probability matrix E=(eiX)
Basic Ideas • As in speech recognition, use Hidden Markov Models (HMM) to model a family of related primary sequences. • As in speech recognition, in general use a left to right HMM: once the system leaves a state it can never reenter it. The basic architecture consists of a main backbone chain of main states, and two side chains of insert and delete states. • The parameters of the model are the transition and emission probabilities. These parameters are adjusted during training from examples. • After learning, the model can be used in a variety of tasks including: multiple alignments, detection of motifs, classification, data base searches.
HMM APPLICATIONS • MULTIPLE ALIGNMENTS • DATA BASE SEARCHES AND DISCRIMINATION/CLASSIFICATION • STRUCTURAL ANALYSIS AND PATTERN DISCOVERY
Multiple Alignments • No precise definition of what a good alignment is (low entropy, detection of motifs). • The multiple alignment problem is NP complete (finding longest subsequence). • Pairwise alignment can be solved efficiently by dynamic programming in O(N2) steps. • For K sequences of average length N, dynamic programming scales like O(NK), exponentially in the number of sequences. • Problem of variable scores and gap penalties.
HMMs of Protein Families • Globins • Immunoglobulins • Kinases • G-Protein-Coupled Receptors • Pfam is a data base of protein domains
HMMs of DNA • coding/non-coding regions (E. Coli) • exons/introns/acceptor sites • promoter regions • gene finding
IMMUNOGLOBULINS • 294 sequences (V regions) with minimum length 90, average length 117, and maximal length 254 • linear model of length 117 trained with a random subset of 150 sequences
G-PROTEIN-COUPLED RECEPTORS • 145 sequences with minimum length 310, average length 430, and maximal length 764. • Model trained with 143 sequences (3 sequences contained undefined symbols) using Viterbi learning.
SOFTWARE STRUCTURE • OBJECT-ORIENTED LIBRARY FOR MACHINE LEARNING • ENGINE IN C++ • GRAPHICAL USER INTERFACE IN JAVA • RUNS UNDER WINDOWS NT AND UNIX (SOLARIS, IRIX)
INFORMATION • ADDITIONAL INFORMATION, POINTERS, REFERENCES, AND SOFTWARE DOWNLOAD: WWW.NETID.COM