Protein Structure 2
Protein Structure 2. Higher Order Protein Structures. Hierarchy of Protein Structure. Primary sequence - The amino acid sequence of a polypeptide, listed from N-terminus to C-terminus.


Protein Structure 2
E N D
Presentation Transcript
Protein Structure 2 Higher Order Protein Structures
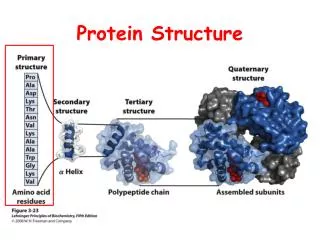
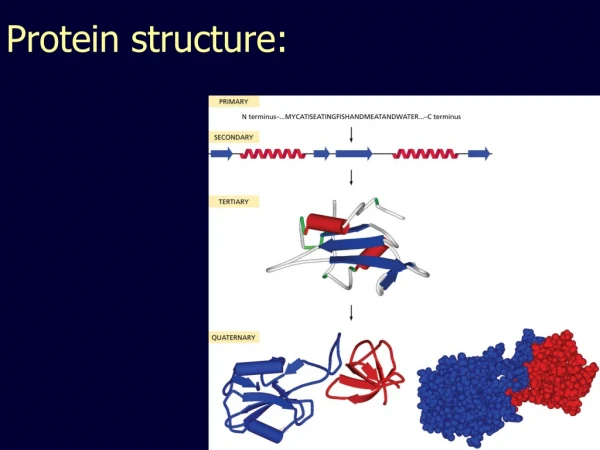
Hierarchy of Protein Structure Primary sequence- The amino acid sequence of a polypeptide, listed from N-terminus to C-terminus. Secondary structure- Recurring structural feature of proteins stabilized exclusively by hydrogen bonds between peptide bond elements. Supersecondary structure- Recurring structural feature of proteins composed of two or more secondary structural elements. Domain- A segment of protein structure that is autonomously stable. Tertiary structure- A stable, independent protein encoded by a single gene. Quaternary structure- A complex structure composed of two or more tertiary structure subunits.
Alpha-fibrous proteins: Coiled-coil Helix Supersecondary Structure Heptad repeats in GCN4 Four-helical bundle
Sequence Comparison Reveals Positional Preference Within Heptad Repeats
Parallel and Anti-parallel Beta Sheets anti-parallel parallel
Supersecondary Structures Structure of -grasp fold of ubiquitin showing motif Repeating motifs (n=7) form a Rossmann fold structure
Supersecondary Structures Greek key motif Beta meander motif
Domains and Tertiary Structure Domains- Structurally autologous structures possessing discrete functions that comprises part of the tertiary structure (beads on a string). Domains can be identified by their characteristic sequences. Ex: DNA binding domains, NAD+/NADP+ binding domains, HECT domains, Zn finger domains, etc. Motif- A highly conserved, short amino acid sequence that identifies a discrete function. Ex: Protein kinase phosphorylation motifs, Ca++ binding motifs, etc. Pyruvate kinase domain structure