Cell Cycle
Cell Cycle. Honors Genetics 2005. Cell cycle and Mitosis Tutorial. http://www.biology.arizona.edu/cell_bio/tutorials/cell_cycle/main.html http://www.cellsalive.com/cell_cycle.htm http://nobelprize.org/medicine/educational/2001/

Cell Cycle
E N D
Presentation Transcript
Cell Cycle Honors Genetics 2005
Cell cycle and Mitosis Tutorial • http://www.biology.arizona.edu/cell_bio/tutorials/cell_cycle/main.html • http://www.cellsalive.com/cell_cycle.htm • http://nobelprize.org/medicine/educational/2001/ Play these three animations to help you learn about the cell cycle and control
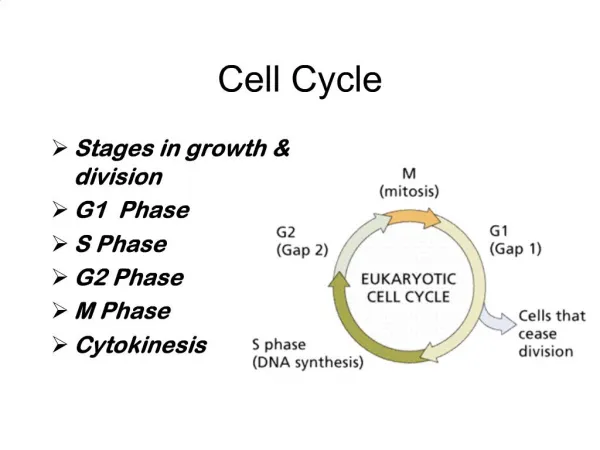
Cell Cycle • G1- Highest metabolic activity. Cytoplasmic contents increase. Protein synthesis, membrane synthesis • S – replication of DNA. Chromosomes copied( sister chromatids form) • G2- Preparation for cell division – Spindle fiber formation • M – Cell division • Cytokinesis – division of the cytoplasm
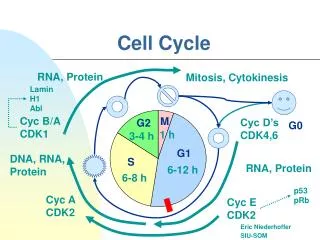
Go • Many times a cell will leave the cell cycle, temporarily or permanently. It exits the cycle at G1 and enters a stage designated G0 (G zero). A G0 cell is often called "quiescent", but that is probably more a reflection of the interests of the scientists studying the cell cycle than the cell itself. Many G0 cells are anything but quiescent. They are busy carrying out their functions in the organism. e.g., secretion, attacking pathogens. • Often G0 cells are terminally differentiated: they will never reenter the cell cycle but instead will carry out their function in the organism until they die.
The passage of a cell through the cell cycle is controlled by proteins in the cytoplasm. Among the main players in animal cells are: Cyclins a G1 cyclin (cyclin D) S-phase cyclins (cyclins E and A) mitotic cyclins (cyclins B and A) Cyclins and cell cycle regulation
Their levels depend upon the stage of the cell cycle They add phosphates to cyclins during the phases of the cell cycle Cyclin-dependent kinases (Cdks) a G1 Cdk (Cdk4) an S-phase Cdk ((Cdk2) an M-phase Cdk (Cdk1) Cyclind dependent kinases
Cyclins - events • A rising level of G1-cyclins bind to their Cdks and signal the cell to prepare the chromosomes for replication. • A rising level of S-phase promoting factor (SPF) — which includes cyclin A bound to Cdk2 — enters the nucleus and prepares the cell to duplicate its DNA (and its centrosomes). • As DNA replication continues, cyclin E is destroyed, and the level of mitotic cyclins begins to rise (in G2).
Checkpoints • DNA damage checkpoints. These sense DNA damage • Before the cell enters S phase (a G1 checkpoint); • During S phase, and • After DNA replication (a G2 checkpoint).
Oncogenes • All the checkpoints examined require the services of a complex of proteins. Mutations in the genes encoding some of these have been associated with cancer; that is, they are oncogenes. This should not be surprising since checkpoint failures allow the cell to continue dividing despite damage to its integrity.
Spindle fibers have three destinations: • Some attach to one kinetochore of a dyad with those growing from the opposite centrosome binding to the other kinetochore of that dyad. • Some bind to the arms of the chromosomes. • Still others continue growing from the two centrosomes until they extend between each other in a region of overlap.
Microtubules • Grow at each end by the polymerization of tubulin dimers (powered by the hydrolysis of GTP), and • Shrink at each end by the release of tubulin dimers (depolymerization)
All types of spindle fibers participate in • the assembly of the chromosomes at the metaphase plate at metaphase. Proposed mechanism (the diagram shows only 1 and 2): • Microtubules attached to opposite sides of the dyad shrink or grow until they are of equal length. • Microtubules motors attached to the kinetochores move them • toward the minus end of shrinking microtubules (a dynein); • toward the plus end of lengthening microtubules (a kinesin). • The chromosome arms use a different kinesin to move to the metaphase plate.
Microtubule Motors • There are two major groups of microtubule motors: • kinesins (most of these move toward the plus end of the microtubules) and • dyneins (which move toward the minus end).
Kinesins and dyneins • The sister kinetochores separate and, carrying their attached chromatid, • move along the microtubules powered by minus-end motors, dyneins, while the microtubules themselves shorten (probably at both ends). • The overlapping spindle fibers move past each other (pushing the poles farther apart) powered by plus-end motors, the "bipolar" kinesins. • In this way the sister chromatids end up at opposite poles.
Mitosis/Meiosis promoting factor • M-phase promoting factor (the complex of mitotic cyclins with the M-phase Cdk) initiates • assembly of the mitotic spindle • breakdown of the nuclear envelope • condensation of the chromosomes • These events take the cell to metaphase of mitosis. • At this point, the M-phase promoting factor activates the anaphase-promoting complex (APC/C) which • Allows the sister chromatids at the metaphase plate to separate and move to the poles (= anaphase), completing mitosis; • Destroys cyclin B. It does this by attaching it to the protein ubiquitin which targets it for destruction by proteasomes.
Anaphase promoting complex • The anaphase-promoting complex (APC). (The APC is also called the cyclosome, and the complex is often designated as the APC/C.) The APC/C • Triggers the events leading to destruction of the cohesins thus allowing the sister chromatids to separate; • Degrades the mitotic cyclin B.
Proteosome – Core Particle • The core particle is made of 2 copies of each of 14 different proteins. • These are assembled in groups of 7 forming a ring. • The 4 rings are stacked on each other (like 4 donuts)
Proteosome – Regulatory Particle • There are two identical RPs, one at each end of the core particle. • Each is made of 14 different proteins (none of them the same as those in the CP). • 6 of these are ATPases. • Some of the subunits have sites that recognize the small protein ubiquitin.
Ubiquitin • A small protein (76 amino acids) • Conserved throughout all the kingdoms of life; that is, virtually identical in sequence whether in bacteria, yeast, or mammals. • Used by all these creatures to target proteins for destruction.
Proteosome and proteins • Are conjugated to a molecule of ubiquitin which binds to the terminal amino group of a lysine residue. • Additional molecules of ubiquitin bind to the first forming a chain. • The complex binds to ubiquitin-recognizing site(s) on the regulatory particle. • The protein is unfolded by the ATPases using the energy of ATP • The unfolded protein is translocated into the central cavity of the core particle.
Proteosomes and Particles(2) • Several active sites on the inner surface of the two middle "donuts" break various specific peptide bonds of the chain. • This produces a set of peptides averaging about 8 amino acids long. • These leave the core particle by an unknown route where • they may be further broken down into individual amino acids by peptidases in the cytosol or • in mammals, they may be incorporated in a class I histocompatibility molecule to be presented to the immune system as a potential antigen [see below]. • The regulatory particle releases the ubiquitins for reuse.
Spindle Checkpoints • Spindle checkpoints. Some of these that have been discovered • Detect any failure of spindle fibers to attach to kinetochores and arrest the cell in metaphase (M checkpoint — example); • Detect improper alignment of the spindle itself and block cytokinesis;
Mitosis occurs in somatic cells • http://www.johnkyrk.com/mitosis.html Check this animation
Mitosis - Prophase • The two centrosomes of the cell, each with its pair of centrioles, move to opposite "poles" of the cell. • The mitotic spindle forms. This is an array of spindle fibers, each containing ~20 microtubules. Microtubules are synthesized from tubulin monomers in the cytoplasm and grow out from each centrosome. • The chromosomes become shorter and more compact.
Mitosis - Prometaphase • The nuclear envelope disintegrates because of the dissolution of the lamins that stabilize its inner membrane. • A protein structure, the kinetochore, appears at the centromere of each chromatid. • With the breakdown of the nuclear envelope, spindle fibers attach to the kinetochores as well as to the arms of the chromosomes. • For each dyad, one of the kinetochores is attached to one pole, the second (or sister) chromatid to the opposite pole. Failure of a kinetochore to become attached to a spindle fiber interrupts the process.
Metaphase • At metaphase all the dyads have reached an equilibrium position midway between the poles called the metaphase plate. The chromosomes are at their most compact at this time.
Anaphase • The sister kinetochores suddenly separate and each moves to its respective pole dragging its attached chromatid (chromosome) behind it.
Telophase • The chromosomes reach the poles • A nuclear envelope reforms around each cluster of • These return to their more extended form.
Cytokineses • In animal cells, a belt of actin filaments forms around the perimeter of the cell, midway between the poles. The interaction of actin and a myosin (not the one found in skeletal muscle) tightens the belt, and the cell is pinched into two daughter cells. • In plant cells, a membrane-bounded cell plate forms where the metaphase plate had been. The cell plate, which is synthesized by the Golgi apparatus, supplies the plasma membrane that will separate the two daughter cells. Synthesis of a new cell wall between the daughter cells also occurs at the cell plate.
Telomeres and aging • http://runews.rockefeller.edu/index.php?page=engine&id=138
....TTGGGGTTGGGGTTGGGGTTGGGGTTGGGGTTGGGGTTGGGGTT 3'....AACCCCAACCCCAACCCC 5' The extra G-T rich sequences are not present in telomeres as single stranded DNA. G-T rich strands can, in the test tube, form an unusual structure in which 4 G's are paired with one another. A metal ion (M+) assists the formation of these G quartets. They stack on top of one another. The T-stretches connect the sides of the quartet. Intermolecular G quartets can form.
Telomeric facts • At their ends, telomeres can assume special structures, the G quartets. • G quartets can account for the stability properties of chromosome ends. • The formation of intermolecular G quartets can account for the apparent interaction of telomeres (bouqet formation). • The studies using TRF2 suggest that the single strand telomeric end is not arranged as a doubled-back G-quartet, but rather invades duplex DNA near the junction of telomeric and non-telomeric DNA. • The analysis of psoralen-treated nuclei suggests that the TRF2-facilitated structures probably exist in actual cells. There is no analogous evidence favoring a biological relevance of G-quartet structures.