Molecular Cloning
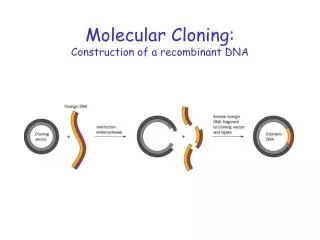
Molecular Cloning. Biology 20L Spring 2003. Overview of Molecular Cloning. Restriction digest of plasmid pUC19 and phage GOAL: Linear pUC19 DNA and several fragments of phage DNA Ligation reaction GOAL: pUC19 recombined with one or more fragments Transformation reaction


Molecular Cloning
E N D
Presentation Transcript
Molecular Cloning Biology 20L Spring 2003
Overview of Molecular Cloning • Restriction digest of plasmid pUC19 and phage • GOAL: Linear pUC19 DNA and several fragments of phage DNA • Ligation reaction • GOAL: pUC19 recombined with one or more fragments • Transformation reaction • GOAL: Use a bacterial host create multiple copies of our new DNA construct (cloning)
Overview of Molecular Cloning • Identification of candidates • GOAL: Determine which bacteria have the desired product (plasmid pUC19 with fragment insert) • Re-isolation of plasmid DNA • GOAL: Obtain multiple copies of the desired DNA construct • Restriction map of plasmid DNA • GOAL: Determine which fragment was inserted in pUC19
Restriction Digest of plasmid pUC19 and phage • Use the restriction enzyme Hind III to cut both plasmid pUC19 and phage . • The recognition sequence for Hind III is: A•AGCTT. This specific sequence occurs once in pUC19, and occurs seven times in phage . • One site in pUC19 creates one linear piece of DNA when cut. • Seven sites in create 8 fragments when cut. • The DNA fragments will be separated and analyzed with gel electrophoresis.
•BstEII pUC19 pUC19 •HindIII •HindIII •BstEII
pUC19 a genetically engineered plasmid • 2.7 Kb (small size allows lots of room for inserting DNA) • Circular non-genomic DNA. • Phage DNA is linear and much larger (48.5 Kb). • Has an origin of replication, and a high copy number. (200+/cell) • Ampicillin resistance gene • Codes for an enzyme that binds and degrades ampicillin. • Lac Z gene (part of the Lac operon) • Codes for ß-galactosidase • An enzyme that breaks down lactose into glucose and galactose. • Polylinker cloning site within LacZ • Contains recognition sequences for several restriction enzymes. • A disruption at this site prevents the production of ß-galactosidase.
http://www.fermentas.com/techinfo/NucleicAcids/mappuc1819.htmhttp://www.fermentas.com/techinfo/NucleicAcids/mappuc1819.htm
T4 Phage T4 phage infecting a bacterial cell
Ligation Reaction • Hind III breaks covalent bonds at the recognition sequence. • A•AGCTT A AGCTT • T TCGA•A TTCGA A • The complimentary or “sticky” ends can readily form H -bonds. • During the ligation reaction, the linearized plasmid and fragments are combined. • Most of these DNA fragments have “sticky” ends. However, because is originally linear, the fragments cut from each end do not have overhangs complementary to the HindIII cuts. • When combined, the pieces form H-bonds in various configurations. • The enzyme DNA ligase is used to form new covalent bonds. • ATP drives this reaction.
Some possible ligation reaction products: Recombinant No insert Fragments No ligation
Transformation • Using bacterial cells to amplify the DNA of interest. • Competent cells are able to take up foreign DNA and acquire genetic information. • Ordinary Escherichia coli cells can be made competent through a treatment with Ca2+ . • Competent cells have very fragile cell walls, and must be handled gently. • During the transformation reaction: • Competent cells are combined with the ligation products. • Incubated on ice (DNA sticks to the outer cell walls.) • Heat Shocked (Membranes become more porous and allow DNA to enter.) • Not all the competent cells will take up DNA. We will determine the frequency. • Incubated in LB broth at 37ºC for ≈ 45 min. • Long enough to allow transcription and translation of ampicillin resistance gene.
Some possible products of the transformation reaction: Bacterial cell Genomic DNA Plasmid w/ insert Ampicillin resistant Nonfunctional LacZ Plasmid w/o insert Ampicillin resistant Functional LacZ No plasmid No ampicillin resistance No LacZ gene Non circular DNA gets degraded within the cells.
Candidate Identification • The transformation culture is plated on special media to help identify which cells have received the recombinant plasmid. • Two types of media: LB + X-gal, & LB+ X-gal + amp • Selection: • Cells with the plasmid can grow on ampicillin media. • Cells without the plasmid cannot grow on ampicillin media. • Screening: • Cells with a functional LacZ gene can convert X-gal to X + gal. • X-gal -------------------------------> X + galactose • Colorless ß-galactosidaseBlue • Cells which produce ß-galactosidase form BLUE colonies. • Cells which are able to grow on ampicillin without ß-galactosidase production form WHITE colonies. (Suspect fragment insert)
Some possible products of the transformation reaction: Bacterial cell Genomic DNA Plasmid w/ insert Ampicillin resistant Nonfunctional LacZ White colony on LB+X-gal+amp. Plasmid w/o insert Ampicillin resistant Functional LacZ Blue colony on LB+X-gal+amp. No plasmid No ampicillin resistance No LacZ gene No growth on Ampicillin
Isolating plasmid DNA • Transformed cells are grown in LB + ampicillin to amplify the target DNA • Selective pressure is important. • E. coli has no instructions for passing the plasmid to the next generation during cell division. • The high number of plasmids within each cell slows growth and lowers the ability to compete. • Plasmid DNA is amplified in two ways: • Cell division (Cells multiply in culture) • High copy replication (multiple copies per cell) • A plasmid miniprep will be performed on cell cultures to extract and purify plasmid DNA. • Cells are disrupted chemically, and the plasmid DNA is separated from genomic DNA and cellular debris.
Final restriction digest • Plasmid DNA isolated from transformed cells will be digested with Hind III, and compared to a known • Hind III marker. • Digests will be analyzed with gel electrophoresis for identification of cloned inserts.
Transformation success • Frequency of transformation = # Transformed cells Total # of cells in the culture • Transformation efficiency = # Transformed cells Amount of DNA in g