Chapt. 4 Molecular Cloning Methods
Chapt. 4 Molecular Cloning Methods. Learning outcomes : Explain major tools of recombinant DNA technology – date from 1970s Expand understanding of new tools Impt. Figs: 1, 2, 3, 4, 5, 10, 11, 12, 15*, 16, 17, 18, 20, 23, 24, 25, 26; Table 1 Review Q: 2, 4*, 6, 7, 8, 9*,

Chapt. 4 Molecular Cloning Methods
E N D
Presentation Transcript
Chapt. 4 Molecular Cloning Methods Learning outcomes: • Explain major tools of recombinant DNA technology – date from 1970s • Expand understanding of new tools • Impt. Figs: 1, 2, 3, 4, 5, 10, 11, 12, 15*, 16, 17, 18, 20, 23, 24, 25, 26; Table 1 • Review Q: 2, 4*, 6, 7, 8, 9*, 10, 11, 13*, 14*; AQ 1
4.1 Gene Cloning in Bacteria • Link prokaryotic or eukaryotic genes to small bacterial DNAs • (plasmids or phage) • Insert recombinant molecules into bacterial hosts to replicate • Can produce large quantities of these genes (or proteins) in pure form Fig. 4
Type II Restriction Endonucleases • Discovered late ‘60s • Named for organism • Recognize specific DNA sequence, cut ONLY at that sequence • Recognize 4-bp to 8-bp sequences • Frequency of cuts less when recognition sequence is longer: (1/4)n • (size of piece larger ~4n)
Restriction Endonucleases, DNA Ligase • Many make staggered cuts in DNA strands • Single-stranded overhangs, sticky ends, can base-pair together (5’-PO4, 3’-OH ends) • Makes joining 2 different DNA molecules easier • DNA ligase needs 5-PO4, 3’-OH ends + ATP • Recognition sequence usually symmetrical palindromes EcoRI: 5’-GAATTC-3’ 3’-CTTAAG-5’
Restriction-Modification System • Enzymes cut foreign DNA, protect cells • What prevents enzymes from cutting up host DNA? • paired methylases recognize, methylate, same site • Form restriction-modification system, R-M system • Methylation protects DNA; after replication, parental strand is partially methylated Fig. 1
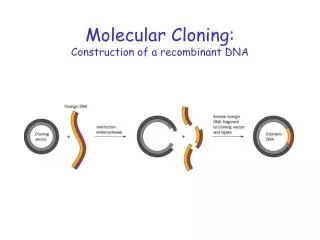
Early Gene Cloning – recombinant DNA • Boyer & Cohen used EcoRI to cut 2 plasmids, small circular pieces of DNA independent of host chromosome • Each plasmid had site for EcoRI • Linear DNA with same sticky ends • Sticky ends base pair • Some ends re-close • Others join 2 pieces • DNA ligase joins 2 pieces with covalent bonds • Transform E. coli; select drugr Fig. 2
Vectors • Vectors function as DNA carriers, permit replication of recombinant DNAs • Typically 1 vector, 1 piece of foreign DNA • Need vector for replication • Foreign DNA has no origin of replication, • (site where DNA replication begins) • Prokaryotic vectors: • Plasmids; Phages • Selectable marker, • Restriction sites for cloning • Origin of replication pBR322
pBR322 Cloning Clone foreign DNA into PstI site of pBR322 • Cut vector to generate sticky ends with PstI • Cut foreign DNA with PstI • Combine vector and foreign DNA: DNA ligase seals • Transform E. coli • (CaCl2, electroporate) • Select Tetr Fig. 4
Screening Transformants pBR322 • Transformation yields bacteria : • Religated plasmid • Religated insert • Recombinants • Identify transformants, recombinants with antibiotic resistance • In tetracycline, only cells with plasmid grow, not just foreign DNA pieces • Test copies of original colonies with ampicillin (kills cells with plasmid including foreign DNA) • Identify desired recombinants Fig. 4
Screen With Replica Plating • Transfer copies of clones from original tetracycline plate to plate with ampicillin • Transfer with sterile velvet • Desired recombinant colonies do NOT grow on ampicillin plate • Pick clone from original plate Fig. 5
pUC plasmid has b-galactosidase fragment • Ampicillin resistance gene • MCS in lacZ’ (N-terminal a-peptide of b-galactosidase) • Foreign DNA in MCS disrupts ability to make b-gal a-peptide: • On X-gal media: clones producing functional b-gal enzyme (from a and host b peptide) produce blue colonies; clones with insert are white. Fig. 6 Fig. 6
Summary Basic Plasmid Cloning Vectors • pBR322 and the pUC plasmids • pBR322 has • 2 antibiotic resistance genes • Many unique restriction sites for inserting foreign DNA • Most of these sites interrupt antibiotic resistance, making screening straightforward • pUC has • Ampicillin resistance gene • MCS that interrupts the b-galactosidase gene a-peptide • MCS also facilitates directional cloning, using pieces cut with 2 different restriction enzymes
Phages As Vectors • Bacteriophages: natural vectors transduce bacterial DNA from one cell to another • Phage vectors infect cells much more efficiently than plasmids transform cells • Clones are not colonies of cells, but rather plaques, clearing of bacterial lawn due to phage infect, kill bacteria in area lambda
Lambda (l) Phage Vectors • Replacement vectors: • Removed middle region (needed only for lysogeny) • Retained genes for phage replication • Replace removed phage genes with foreign DNA • Phage vectors hold larger amounts of foreign DNA • Accept up to 20-kb of DNA • Traditional plasmid vectors usually less • Lambda phage vectors require minimum size DNA (12 kb) insert to package into phage particle
Cloning Using a Lambda Phage Vector • Join DNA in vitro • Add packaging • proteins from cell • extract • Infect cells Fig. 8
Genomic Libraries • Large number of clones: all genes from species’ genome • Restriction fragments can be packaged into phage, about 16 – 20 kb per fragment (often partial digests) • Need thousands of clones for library: ex. Yeast genome 107 bp/ 104 bp/clone -> ____ clones ex. Human genome 3 x 109 bp /104 bp/clone -> ___ • Once library is established, search for gene of interest (BL 415 experiment with plasmid library)
Plaque Hybridization • Search genomic library with probe to identify clone containing desired gene • Ideal probe – labeled nucleic acid with sequence matching the gene of interest Fig. 9
Cosmids Cosmids designed to clone large DNA fragments • Behave as plasmid and phage • Contain • cos sites, cohesive ends of l phage DNA, allow DNA to be packaged into l phage head • Plasmid origin of replication permits replication as a plasmid in bacteria • Most l genome removed so hold large inserts (40-50 kb) • So little phage DNA, can’t replicate like phage; infectious, carry recombinant DNA into bacterial cell, then plasmid
M13 Phage Vectors • Long, thin, filamentous phage M13 • Normal size 6.4 kb (10 genes) • Modified vector contains: • Gene fragment with b-galactosidase • Multiple cloning site like pUC plasmid • Advantages • Can hold large DNA; length of phage increases • Phage genome is single-stranded DNA • Fragments cloned (into ds intracellular form) are recovered in single-stranded form • Phage does not kill cells; extrudes particles • Useful for sequencing DNA, mutagenesis
M13 Cloning permits recovery of Single-stranded DNA • After infecting E. coli, single-stranded phage DNA was converted to double-stranded replicative form (RF, plasmid-like) • Purify replicative form to clone foreign DNA into MCS; ligate vitro • Transform recombinant DNA into host cells, results in plasmid-like RF molecule in cells • Phage particles, contain one single-stranded DNA (+ strand), secreted from transformed cells, can be collected from media Fig. 10
Phagemids are versatile Vectors: pBluescript(pBSK) Aspects of both phages and plasmids: • Ampr for selection of clones • MCS inserted into lacZ’ gene permit blue/white screening • Origin of replication of single-stranded phage f1 permits recovery of single-stranded recombinant DNA (like M13) • MCS has phage RNA polymerase promoters, 1 on each side of MCS; permits specific RNA transcripts in vitro Fig. 11
Eukaryotic Vectors,High Capacity vectors • Plasmids and Viruses clone genes in eukaryotic cells • Plasmid shuttle vectors (origins of replication for both bacterial and eukaryotic host cell; ex. Yeast YEP24 • Selectable markers in both hosts; MCS sites • Yeast artificial chromosomes (YAC), bacterial artificial chromosomes (BAC) clone huge pieces • Vectors based on Ti plasmid for plant cells
Identifying specific clone with specific probe • Probes can identify a desired clone from among thousands of irrelevant ones: • Two types widely used: Recognize gene or mRNA with Oligonucleotides: Short RNA or DNA sequences that– hybridize • Can use homologous gene from another organism • If amino acid sequence is known, deduce possible nucleotide sequences to code for these amino acids Recognize specific protein product with Antibodies
cDNA Cloning from Eukaryotes into Bacteria • cDNA: complementary DNA or copy DNA • made from mRNA • A cDNA library: set of clones representing many mRNAs from cell type or tissue or time • Library can contain thousands of different clones • * Need cDNA clones because genomic clones have introns, aren’t expressed in bacteria
Making cDNA Library from eukaryotic mRNA • Reverse Transcriptase (RT) • mRNA -> DNA • Primer • oligo(dT) to polyA at 3’ end • RNAse H degrades RNA • DNA polymerase Pol I finishes • Terminal transferaseto add C tails and G tails for cloning Fig. 12
Vector choice for cDNA cloning • Purpose and method to detect positive clones • If cloning gene and small fragment, plasmid or phagemid like pUC or pBS used, with colony hybridization and labeled DNA probe • If large fragments, use cosmids, or l phage or YAC, BAC • For expression of gene product, use cDNA of eukaryote cloned in pUC, phagemids of lambda • under control of lac promoter for transcription and translation of gene; use hybridization, antibody screening (more later)
Summary clone cDNA into Bacteria • Make cDNA library with synthesis of cDNAs one strand at a time • Use mRNAs from a cell as templates for 1st strands, then 1st strand as template for 2nd • Reverse transcriptase generates 1st strand • DNA polymerase I generates 2nd strand • Give cDNAs oligonucleotide tails that base-pair with complementary tails on a cloning vector • Use recombinant DNAs to transform bacteria • Detect clones with: • Colony hybridization using labeled probes • Antibodies if gene product is translated
Polymerase chain reaction(PCR): amplifies DNA exponentially;yields DNA fragments for cloning, forensics, diagnostics Fig. 15
Standard PCR • Invented by Kary Mullis in 1980s • DNA polymerase copies selected region of DNA • Short DNA primers hybridize to DNA on either side of piece of interest – initiate DNA synthesis through that area, X • Selected region DNA doubles in amount each cycle • Special heat-stable polymerases work after high temperatures that separate (denature) strands simplifies process – ex. Taq polymerase • Real-time PCR uses fluorescent dyes to quantify amplification, reflect amount of mRNA or DNA in cells • (Fig. 17)
Reverse transcriptase PCR (RT-PCR) to clone specific cDNA Start with mRNA, not dsDNA Convert mRNA to DNA with primer specific to gene Use forward primer to convert ssDNA to dsDNA Then standard PCR • With primer choice, restriction enzyme sites added to cDNA • 2 sticky ends permit directional cloning – good for protein expression Fig. 16
4.3 Expressing Cloned Genes • Cloned gene permits production of large quantities of particular DNA sequence • Expression vectors yield protein products • Strong promoter, ribosome binding site near ATG codon • Make as pure protein, or as fusion to bacterial peptide • Regulated expression: Inducible expression vectors keep cloned gene repressed until time to express: • Large quantities of eukaryotic protein can be toxic
Fusion proteins:Oligo-histidine(His-tagged) Expression Vector • Sequence upstream of MCS encodes 6 His which get added to protein: • Oligo-histidine high affinity for divalent metal ions (Ni2+) • Purify protein by nickel affinity chromatography • His tag removed using enzyme enterokinase if needed Fig. 20
Fusion Proteins with lacZ’ • Cloning vectors (pUC and pBS) are expression vectors using lac promoter with lacZ’a-peptide • If inserted DNA in same reading frame as interrupted gene, fusion protein results • Partial b-galactosidase at N- end • Inserted protein at C-end • Fusions often more stable • Fusions also assist detection, purification Fig. 18
Regulation: lac Promoter/ operator • lac promoter inducible with lactose or synthetic IPTG; binds repressor and removes it from O • Transcription stays off until induced
Fusion Proteins also in lgt11 vector • Phage contains lac control region and lacZ’ gene (a-peptide) • Products of gene correctly inserted are fusion proteins with b-galactosidase peptide • Use blue-white screening on host with b-peptide • Fusion reduces degradation of foreign protein Fig. 21
Antibody screening with lgt11 (or plasmids) • Lambda phages with cDNA inserts on E. coli • Proteins released are blotted onto membrane • Probe with specific 1oantibody to protein • Antibody bound to protein from plaque is detected with labeled 2o antibody Fig. 22
Summary – Expression in Bacteria • Expression vectors can produce fusion proteins • One part from coding sequences in vector • Other part from sequences in cloned gene • Many fusion proteins can be simply purified by affinity chromatography (His-tag; Flag-tag; TAP-tag) • Fusion proteins can be detected with antibodies after blotting of colonies or plaques:
Bacterial Expression System Shortcomings • Bacteria recognize proteins as foreign and destroy • Posttranslational modifications (glycosylation, phosphorylation) are not done correctly • Bacterial cytoplasm not permit correct protein folding • High levels of cloned eukaryotic proteins can be expressed in useless, insoluble (inclusion) form
Use Eukaryotic Expression Systems • Initial cloning in E. coli using shuttle vector, that replicates in both bacterial and eukaryotic cells • Ex. Yeast Saccharomycescerevisiaewell-suited : • Rapid growth, ease of culture • Multicopy plasmids (from natural 2 m plasmids) • Eukaryote, so more appropriate posttranslational modifications to expressed proteins • Can secrete protein for easy purification • Ex. Plasmid YEP24 shown earlier
Ex. Baculovirus expression vectors;transform (transfect) insect cells in culture • Large circular DNA genome; • Major viral structural protein made in huge amounts • Strong promoter for polyhedrin protein • Produce lots of recombinant protein per liter of medium • Non-recombinant viral DNA do not replicate because lack essential gene Fig. 23
Methods for Transfection of animal cells with vectors for clones or protein production [Transformation for animal cells used for cancer cells] • Calcium phosphate • Mix cells with DNA in phosphate buffer • Solution of calcium salt added to form precipitate • Cells take up calcium phosphate crystals, includes DNA • Electroporation • Electric shock for DNA to enter cells • Liposomes • DNA mixed with lipid to form liposomes, small vesicles • Liposomes fuse with cell membrane, carry DNA inside
Summary • Foreign genes can be cloned and expressed in eukaryotic cells • Eukaryotic systems have some advantages: • Proteins tend to fold properly, are soluble rather than aggregated into insoluble inclusion bodies • Posttranslational modifications (phosphorylation, glycosylation) made in eukaryotic manner • Can use transient clones for tests gene function • Select stable clones expressing transgene (as for construction transgenic organisms)
Review problems 2. Why does one need to attach DNAs to vectors to clone them? • Diagram the process of cloning a DNA fragment into BamHI and PstI sites of vector pUC18. How would you screen for clones that contain an insert? Explain the blue/white lacZ’ detection system 7. You want to make a genomic library with DNA fragments averaging about 45 kb in length. Which vector discussed in this chapter would be most appropriate? Why?
Review problems 11. Diagram a method for creating a cDNA library. • Outline the polymerase chain reaction (PCR) method for amplifying a given stretch of DNA. • What is the difference between reverse transcriptase PCR (RT-PCR) and standard PCR? For what purpose would you use RT-PCR? For what purpose might you want cDNAvs genomic library?
Review problems AQ 1. Given an amino acid sequence of part of a protein from a hypothetical gene you want to clone: Pro-arg-tyr-met-cys-trp-ile-leu-met-ser Look at the code chart • What sequence of 5 aa would give a 14-mer probe with the least degeneracy for probing a library to find gene of interest? • How many different 14-mers would you need to make to be sure probe matches sequence in cloned gene perfectly? • lf you started your probe one aa to the left of the one you chose in (a), how many different 14-mers would you have to make?