Enzymes
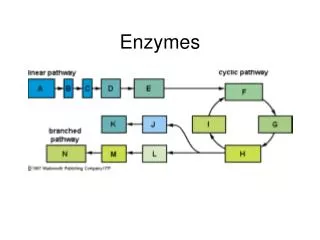
Enzymes. 1. Objectives. What are enzymes ? Properties of enzymes Classification Factors Affecting Enzyme Action Enzyme Kinetics. What Are Enzymes?. Biological Catalyst Most enzymes are Proteins Not permanently changed in the process Not consumed. 4.

Enzymes
E N D
Presentation Transcript
Enzymes 1
Objectives • What are enzymes ? • Properties of enzymes • Classification • Factors Affecting Enzyme Action • Enzyme Kinetics
BiologicalCatalyst Most enzymes are Proteins Not permanently changed in the process Not consumed 4
Differ from inorganic catalysts in: Being sensitive to changes in pH , temperature , loss of activity by time specificity
Definitions • Ribozymes : are RNAs with catalytic activity. • catalyze the cleavage & synthesis of phosphodiesterbonds • Zymogen • Isozymes
Isoenzymes: • enzymes that catalyze the same reaction but may have genetically determineddifference in the amino acid sequence • Thus may have difference in their physical properties
Plasma enzymes 2 major groups: • 1-Functional plasma enzymes • perform specific physiological function. • Examples: • Zymogens (inactive precursors) of enzymes involved in blood coagulation. • 2-Non-functional plasma enzymes • no physiologicalrole in the blood. • occur in blood in very minute amounts).
Enzymes composed wholly of protein are known as simple enzymes • Complex enzymes, which are composed of protein + small molecule holoenzymes. • Protein component apoenzyme, while the non-protein component coenzyme or cofactor
Prosthetic group describes a complex in which the small organic molecule is permanently bound to the apoenzyme by covalent bondsand cosubstrate occurs when it is transiently attached. • Coenzymes are often derivatives of vitamins (FAD,NAD, CoA)
Catalytic efficiency: Typically 100-1000 substrate molecules transformed to product / second ?(by enz.) Turn over number : is the number of substrate moles converted to product/enzyme mol /sec.
Location within the cell: • Mitochondria: TCA cycle Fatty oxidation Decarboxylation • Cytosol: Glycolysis HMP Fatty acid synthesis • Lysosomes: Degradation of macromolcule
Recommended name uricase ,glucosidase
Systematic Name • Currently enzymes are grouped into six functional classes by the International Union of Biochemistry & Molecular Biology (I.U.B.M.B
Regulation: Rate of product formation responds to the needs of the cells
Factors Affecting Enzyme Action
Temperature Optimum temperature Reaction Rate Low High Temperature
Substrate Concentration Maximum activity Reaction Rate substrate concentration
pH: Narrow range of activity Most lose activity in low or high pH Reaction Rate Optimum pH 3 5 7 9 11 pH
Time • This is because by time the substrate decreases (↓v1 ) , • the product increases(↑v-1 ) besides • the enzyme decreases due to denaturation.
How do enzymes Work? Enzymes work by weakening bonds which lowers activation energy 27
Substrate Transition state Product If enzyme just binds substrate then there will be no further reaction X Enzyme not only recognizes substrate, but also induces the formation of transition state
Enzymes Without Enzyme With Enzyme Free Energy Free energy of activation Reactants Products Progress of the reaction 29
The active site consists of certain groups e.g.-SH, -OH, COO-, NH3 + & imidazole. Some groups are concerned with binding of the substrate & that determines the specificity of the enzyme.
Other groups are concerned with catalysis. • In some enzymes these groups can participate in general acid-base catalysis. In others, catalysis may involve transient formation of a covalent enzyme-substrate complex.
Inhibition of enzyme activity • Inhibitor :is any substance that can diminish the velocity of the enzyme catalyzed reaction. • Competitive • Non competitive: Heavy metal ions (e.g. mercury and lead) should generally be prevented from coming into contact with enzymes as they usually cause such irreversible inhibition by binding strongly to the amino acid backbone. • Uncompetitive
Enzyme Inhibition (Mechanism) E + S→ES→E + P + I ↓ EI E + S→ES→E + P + + II ↓ ↓ EI+S→EIS E + S→ES→E + P + I ↓ EIS ← ← ← ↑ ↑ ↑ ↑ Uncompetitive Competitive Non-competitive E Substrate E X Cartoon Guide Compete for active site Inhibitor Different site Equation and Description [I] binds to free [E] only, and competes with [S]; increasing [S] overcomes Inhibition by [I]. [I] binds to [ES] complex only, increasing [S] favors the inhibition by [I]. [I] binds to free [E] or [ES] complex; Increasing [S] can not overcome [I] inhibition. Juang RH (2004) BCbasics
Sulfa Drug Is Competitive Inhibitor -COOH H2N- -SONH2 H2N- Para-aminobenzoic acid (PABA) Bacteria needs PABA for the biosynthesis of folic acid Folic acid Tetrahydro- folic acid Pteridine Precursor Sulfa drugs has similar structure with PABA, and inhibit bacteria growth. Sulfanilamide Sulfa drug (anti-inflammation) Adapted from Bohinski (1987) Modern Concepts in Biochemistry (5e) p.197
Regulation of Enzyme Activity • Substrate availability • Allosteric Effectors • Covalent modification • Induction & repression
Regulation maybe short term regulation or long term regulation • The long term regulation includes enzyme synthesis & repression • The short term regulation includes allosteric regulation& covalent modification
Allosteric Effectors • Allosteric means (“occupy another space”, other than that of the substrate). • Allosteric enzymes are enzymes whose activity at the catalytic site may be modulated by the presence of allosteric effectors at an allosteric site (i.e. at a site other than the active site.) • Allosteric enzymes usually contain multiple subunits. • The molecules regulating the allosteric enzymes are called effectors (modifiers or modulators).
Allosteric effectors bind non-covalentlyat a site other than the active site. • Binding of allosteric effectors at the allosteric site induces conformational changes at the catalytic site& this can alter the affinity of the enzyme for its substrate or modify the maximal catalyticactivity of the enzyme or both.