Phylogenetic Analysis of Rice Blast Resistance Proteins and Related Wild Potato Protein
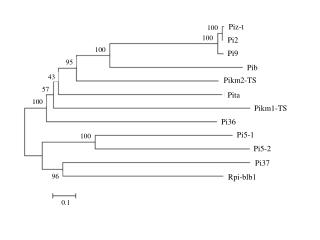
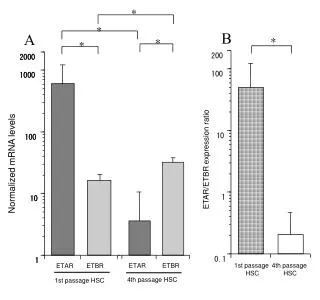
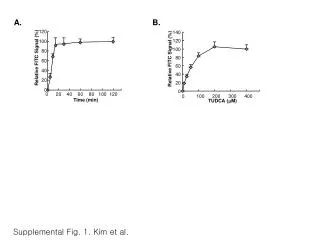
This supplementary figure presents a phylogenetic tree constructed from cloned rice blast resistance proteins, including the wild potato protein Rpi-blb1, which confers resistance to Phytophthora infestans. The tree was generated using alignments of full protein sequences and the neighbor-joining method, with bootstrap values indicating reliability. Each branch percentage is based on 1,000 bootstrap replicates, and the scale bar reflects a distance of 10 changes per 100 amino acid positions. Accession numbers for all analyzed proteins are provided for reference.

Phylogenetic Analysis of Rice Blast Resistance Proteins and Related Wild Potato Protein
E N D
Presentation Transcript
Piz-t 100 100 Pi2 100 Pi9 95 Pib 43 Pikm2-TS 57 Pita 100 Pikm1-TS Pi36 Pi5-1 100 Pi5-2 Pi37 Rpi-blb1 96 0.1
SUPPLEMENTARY FIGURE S3. Phylogenetic tree of cloned rice blast resistance proteins. The wild potato protein Rpi-blb1 that confers resistance to P. infestansand that shows relatively high similarity to Pi5 proteins is included in this analysis. Alignments of the whole protein sequences were used to generate the phylogenetic tree via the neighbor-joining method. Numbers on branches indicate the percentage of 1,000 bootstrap replicates. The scale bar corresponds to a distance of 10 changes per 100 amino acid positions. The accession numbers are as follows: Pi5-1 (EU869185), Pi5-2 (EU869186), Piz-t (ABC73398), Pi2 (ABC94599), Pi9 (ABB88855), Pib (BAA76282), Pita (AAK00132), Pi36 (ABI64281), Pi37 (ABI94578), Pikm1-TS (BAG72135), Pikm2-TS (BAG72136), and Rpi-blb (Q7XBQ9).